Citation: Tonini JFR, Forlani MC, de Sá RO (2014) A new species of Chiasmocleis (Microhylidae, Gastrophryninae) from the Atlantic Forest of Espírito Santo State, Brazil. ZooKeys 428: 109–132. doi: 10.3897/zookeys.428.7352
Among Neotropical microhylids, the genus Chiasmocleis is exceptionally diverse. Most species of Chiasmocleis were described in recent years based on external morphology, but recent studies using molecular data did not support the monophyly of the species groups clustered based on feet webbing. Furthermore, a phylogeographic study of C. lacrimae estimated high genetic divergence and low gene flow among populations across small geographic ranges. Increasing the molecular and geographic sampling, and incorporating morphological data, we identified new cryptic species. Herein, we used novel genetic and morphological data to describe a new species of Chiasmocleis.
Amphibians, Chiasmocleis quilombola sp. n. , cryptic species, phylogenetics, systematics
The diversity of evolutionary lineages with little phenotypic differences (i.e., cryptic species) might be better understood in the light of genetic delimitation of evolutionary units (
Species are segments of population level evolutionary lineages and do not necessarily need to be phenetically distinguishable, diagnosable, monophyletic, intrinsically reproductively isolated, ecologically divergent, or anything else to be considered species, but they only have to be evolving separately from other lineages (
A recent molecular phylogeny (
Herein, we describe a new species of Chiasmocleis from the Atlantic Forest of southeastern Brazil and present a phylogenetic hypothesis for the species group.
Specimens and tissues used herein and comparative material are deposited in the following collections: 1) CFBH: Coleção de Anfíbios Célio Fernando Baptista Haddad, Departamento de Zoologia, Universidade Estadual Paulista Rio Claro, Rio Claro, São Paulo State, Brazil; 2) MNRJ: Museu Nacional do Rio de Janeiro, Rio de Janeiro, Rio de Janeiro State, Brazil; 3) Museu de Zoologia, Universidade de São Paulo, São Paulo, São Paulo State, Brazil; 4) MBML: Museu de Biologia Mello Leitão, Santa Teresa, Espírito Santo State, Brazil; 5) CTA: Coleção de Tecidos e DNA da Universidade Federal do Espírito Santo (UFES) and LGA: Laboratório de Genética Animal, Vitória, Espírito Santo State, Brazil; 6) RN and CTRN: Universidade Federal Rural do Rio de Janeiro, Seropédica, Rio de Janeiro State, Brazil. Field numbers correspond to M. T. Rodrigues (MTR), Universidade de São Paulo, São Paulo, São Paulo State, Brazil; P. Rocha (PEU), Universidade Federal da Bahia, Salvador, Bahia State, Brazil; and J. F. R. Tonini (JFRT), vouchers are at UFES. Specimens examined and tissues samples are listed in Appendix 1 and 2, respectively, and sample localities are shown in Figure 1.
Sample localities of A tissues and B specimens included in the present study. Sites with more than one color indicates syntopy. List of localities: 1 Porto Seguro, 2 Trancoso, 3 ReBio Córrego Veado, 4 FloNa do Rio Preto (type locality of Chiasmocleis quilombola sp. n.), 5 Parque Estadual de Itaúnas, 6 ReBio Sooretama, 7 Reserva Natural Vale, 8 Povoação, 9 FloNa dos Goytacazes, 10 Costa Bela, 11 ReBio Duas Bocas, 12 Guarapari, 13 Mata da Usina Paineiras, 14 Mimoso do Sul, 15 ReBio União, 16 Cachoeiras de Macacu, 17 Duque de Caxias, 18 Angra dos Reis, 19 Picinguaba, 20 Ilha de São Sebastião, 21 Bertioga, 22 Aracruz (type locality of Chiasmocleis capixaba), 23 Horto Florestal (type locality of Chiasmocleis lacrimae). Blue lines represent major coastal rivers, from North to South: Jequitinhonha, Mucuri, Doce, and Paraíba do Sul. BA = Bahia State, ES = Espírito Santo State, RJ = Rio de Janeiro State, SP = São Paulo State, and MG = Minas Gerais State.
The following measurements were adapted from
Molecular Analyses: Total genomic DNA was extracted from ethanol-preserved liver or muscle tissues using Qiagen DNeasy kit (Valencia, California, USA). We used four molecular markers (mtDNA: 12S, 16S, and NADH dehydrogenase subunit 2 [ND2]; nucDNA: brain-derived neurotrophic factor [BDNF]), amplified using previously published primer sets and PCR profiles (
The following outgroup were chosen based on published phylogenies including species of Chiasmocleis (
Best partition scheme and substitution models selected using Partition Finder.
| Subset | Best Model | Subset partitions | Subset sites |
|---|---|---|---|
| 1 | HKY+I+G | 12S | 1–700 |
| 2 | HKY+G | 16S, ND2_1 | 701–1044, 1661–2473\3 |
| 3 | K80+I | BDNF | 1045–1660 |
| 4 | HKY+G | ND2_2 | 1662–2473\3 |
| 5 | GTR+G | ND2_3 | 1663–2473\3 |
We applied two approaches of phylogenetic estimation: 1) Maximum Likelihood (ML) and 2) Bayesian inference (BI) using the dataset containing the markers 12S, 16S, ND2, and BDNF. Maximum Likelihood in RAXML v7.2.8 (
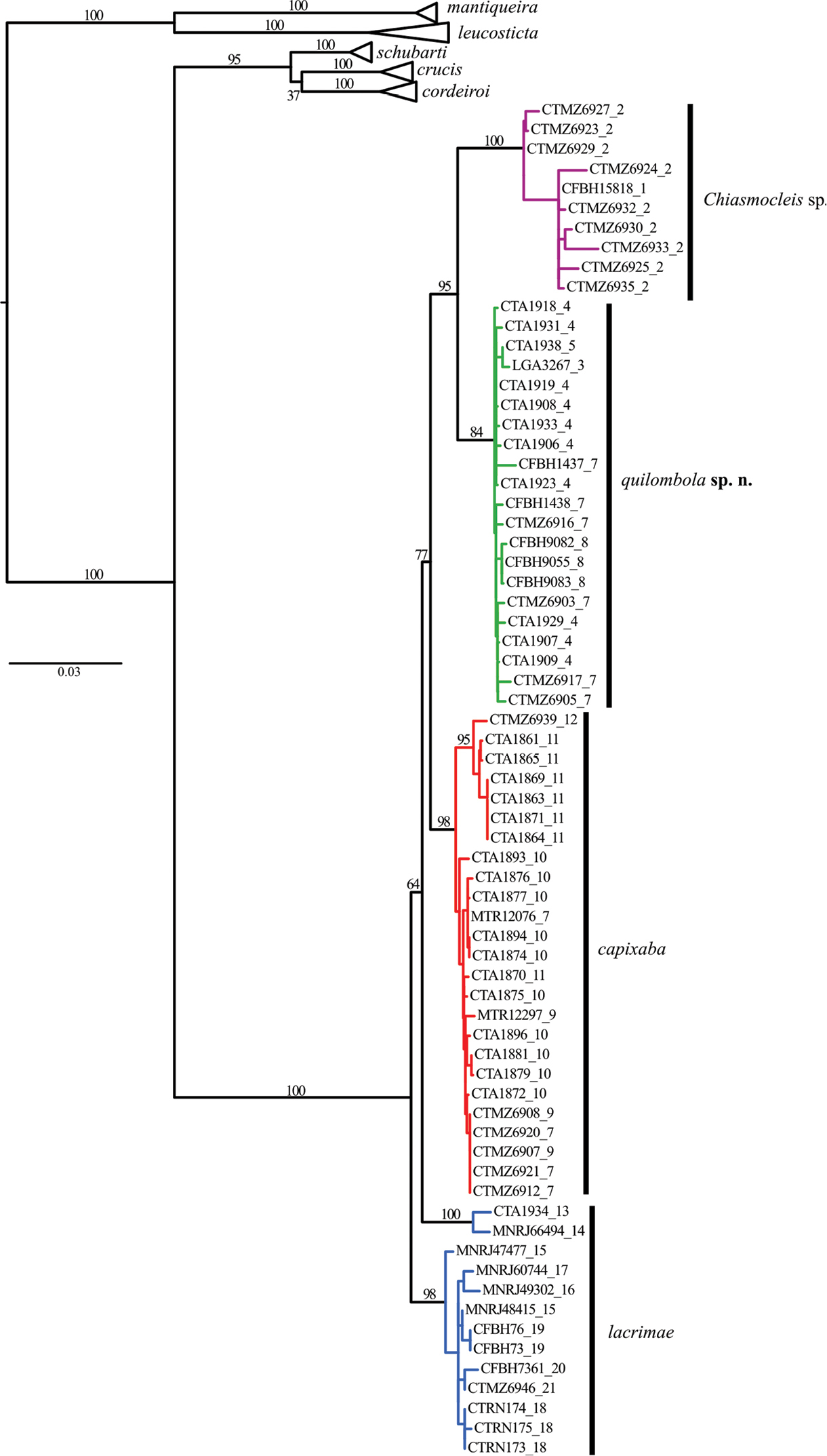
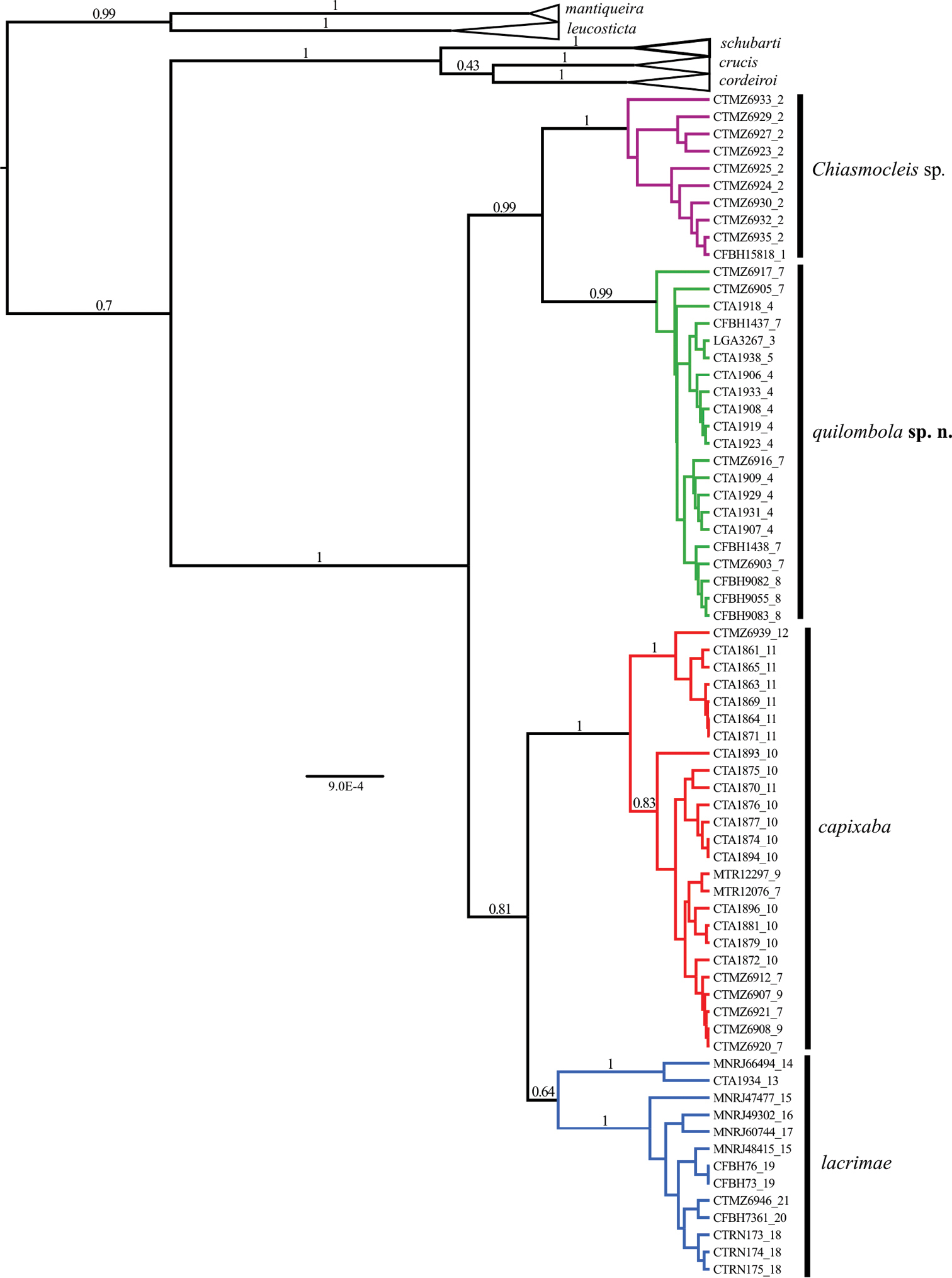
The phylogenetic hypotheses generated through ML (Figure 2) and the BI (Figure 3) resulted in similar topology. The ML tree (Figure 2) supported two new species as sister group of Chiasmocleis capixaba, Chiasmocleis lacrimae correspond to a basal node; whereas in the BI tree (Figure 3) Chiasmocleis capixaba was estimated as sister to Chiasmocleis lacrimae, but the posterior probability of this node was lower than 0.95. Both the ML and BI trees showed clades of Chiasmocleis leucosticta, Chiasmocleis mantiqueira, Chiasmocleis crucis, Chiasmocleis schubarti, and Chiasmocleis cordeiroi, but not Chiasmocleis lacrimae and Chiasmocleis capixaba (Figure 2, 3). Populations of Chiasmocleis capixaba and Chiasmocleis lacrimae from the north of the Espírito Santo State, as well as populations of Chiasmocleis lacrimae from southern areas of the Bahia State, would represent two new cryptic lineages closely related to Chiasmocleis lacrimae and Chiasmocleis capixaba. In the ML analysis populations from southern Espírito Santo formed a clade that makes Chiasmocleis lacrimae polyphyletic (Figure 2), whereas in the BI these populations formed a clade including also populations of Chiasmocleis lacrimae from the states of São Paulo and Rio de Janeiro (Figure 3). However, given the lack of support for this node in both analyses, basing taxonomic change on the presumed polyphyly is not warranted at present.
Maximum likelihood tree including 12S, 16S, ND2, and BDNF. Node numbers correspond to bootstrap, values >70 indicate good support. Although Chiasmocleis lacrimae may not represent a monophyletic species, bootstrap values are low to make further assumptions. Numbers after underscore symbol correspond to localities present in Figure 1. Scale bar represents number of substitutions/site.
Phylogenetic hypothesis obtained through Bayesian Inference using 12S, 16S, ND2, and BDNF. Node numbers correspond to posterior probabilities, values >0.95 indicate good support. Numbers after underscore symbol correspond to localities present in Figure 1. Scale bar represents number of substitutions/site.
Therefore, our results show that the new species clusters within the genus Chiasmocleis.
MZUSP147478, adult male, collected at the Floresta Nacional do Rio Preto (Figure 4A), Municipality of Conceição da Barra, Espírito Santo State, Brazil (18°21'19"S, 39°50'39"W), collected on December 8-16, 2009, by L. P. Costa, J. F. R. Tonini, J. Dalapicolla, R. Duda, and C. M. Mattedi.
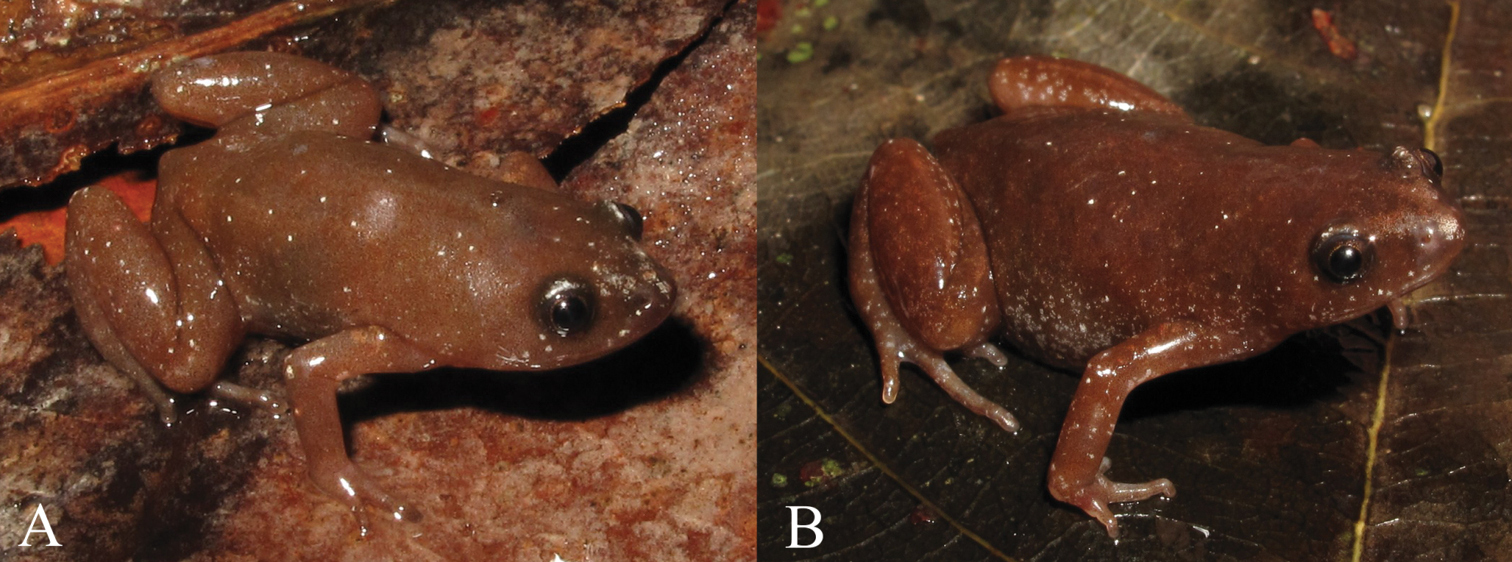
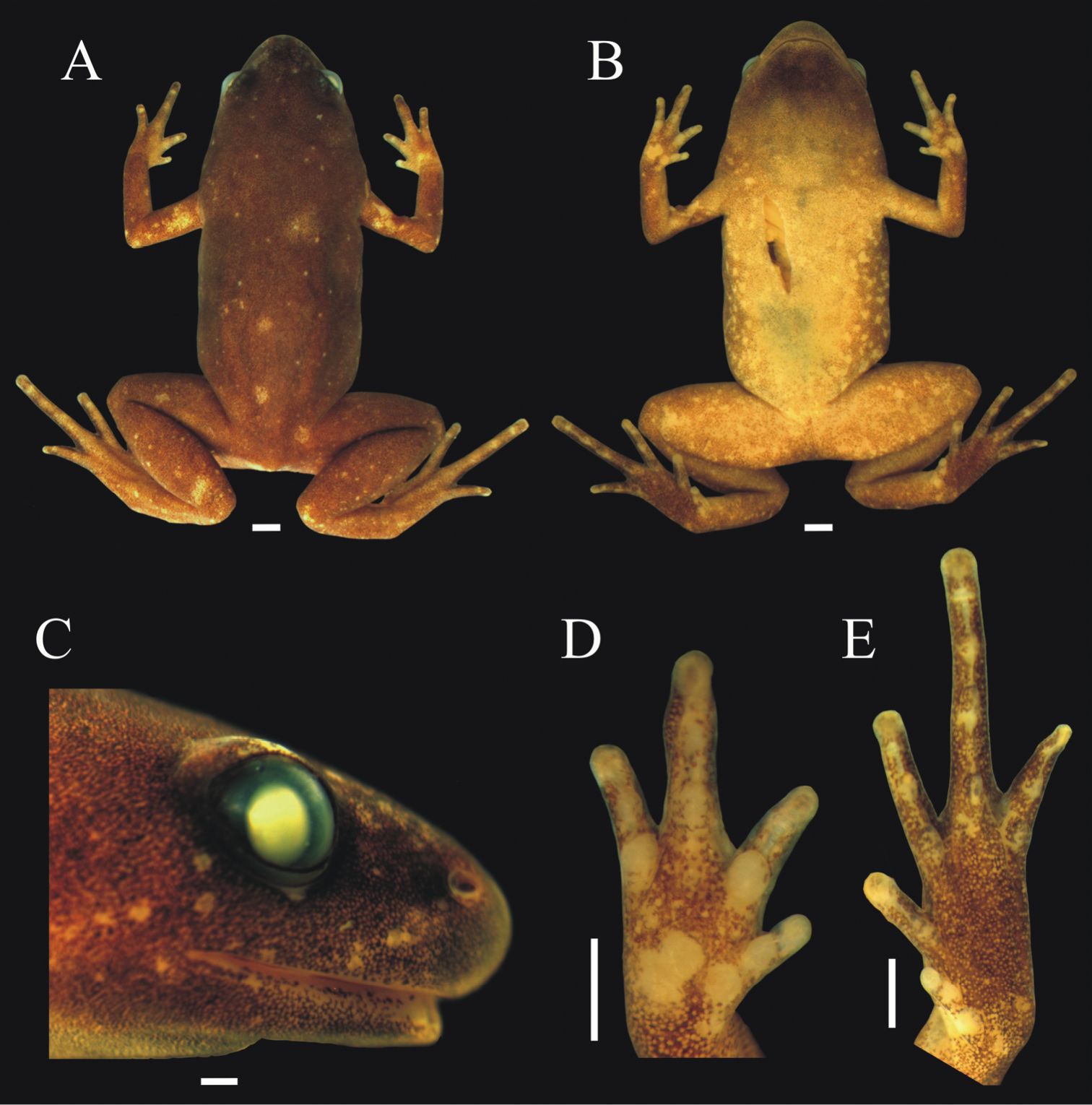
Chiasmocleis quilombola sp. n. in vivo. A male (holotype: MZUSP147478) and B female (MZUSP147479, paratopotype). Not in scale.
Males: MZUSP147471–73, MZUSP147475–76, MZUSP147494; female: MZUSP147479 (Figure 4B), Municipality of Conceição da Barra, Espírito Santo State, Brazil (18°21'19"S, 39°50'39"W), collected on December 8-16, 2009, by L. P. Costa, J. F. R. Tonini, J. Dalapicolla, R. Duda, and C. M. Mattedi.
A small-sized species of Chiasmocleis (males SVL mean= 14 ± 1.4 mm; female SVL = 17.1 mm), diagnosed by the following combination of characters: (1) body slender; (2) snout rounded in lateral and dorsal views; (3) all fingers slightly fringed, not webbed, in males and female; (4) all toes fringed and slightly webbed in males and female; (5) dermal spines on fingers and toes of males can be present or absent, absent in female; (6) dermal spines on dorsal surface of males can be present or absent, absent in female; (7) dermal spines absent on ventral surface in males and female; (8) dermal spines on chin and snout of males can be present or absent, absent in female; (9) dermal spines over outer surfaces of legs and cloaca in males can be present or absent, absent in female; (10) female has para-cloacal glands; (11) incomplete occipital fold; (12) vocal slits present in males; (13) dorsal coloration brown; (14) medial ventral body surface light cream colored, whereas ventrolateral surfaces have a light brown and cream marbled pattern; (15) ventral surfaces of fore and hind limbs with a homogeneously and finely dark pattern over a cream background; (16) dorsal surface of fore and hind limbs light brown with a few cream spots or blotches, more distinct on the fore limbs; (17) male throat infuscate; (18) mid-dorsal and/or line on posterior surface of thighs may be present; and (19) tympanum indistinct.
Body small (SVL = 15.7 mm), slender, slightly ovoid (Figure 5); head triangular in shape, broader than long; snout short, tip of snout rounded (Figure 5A–B); nostrils located closer to the tip of snout than to eye, not protuberant, directed laterally (Figure 5C); inter-nostril distance smaller than eye–nostril distance and smaller than eye diameter; canthus rostralis slightly defined; loreal region slightly convex; lips not flared; eyes small, slightly protruding; inter-orbital area flat; incomplete occipital fold; tympanum indistinct; upper jaw projecting beyond lower one; tongue large, elongate, and laterally free; premaxillae, maxillae, and vomerine teeth absent; choanae small, rounded, widely separated, positioned anterolaterally to eye; vocal slit present.
Holotype of Chiasmocleis quilombola sp. n. (MZUSP 147478). A Dorsal B ventral, and C lateral views D right hand and E right foot. White bars = 1 mm.
Arms slender, lacking tubercles on forearm. Hands not webbed (Figure 5D); fingers tips rounded, not expanded, and slightly fringed; fingers lacking dermal spines; finger lengths I<II<IV<III; thumb without nuptial asperities; subarticular tubercles well developed and rounded, proximal subarticular tubercles larger than others; supernumerary tubercles absent; thenar tubercle well developed, ovoid, and at the base of finger I; two palmar tubercles, a rounded inner tubercle and an elongated outer one (Figure 5D). Legs short, moderately robust; knee and heel lacking tubercles; tibial and tarsal ridges absent. Foot slightly webbed (Figure 5A–B, E); toes slightly fringed; toe tip rounded lacking disks; subarticular tubercles well developed, ovoid; supernumerary tubercles absent; an oval inner, but no outer, metatarsal tubercle. Toe lengths I<II<V<III<IV; toes lacking dermal spines; tibia length slightly shorter than thigh length; combined thigh and tibia lengths approximately 82.8% of snout-vent length; foot length approximately 43.9% of snout-vent length.
Skin smooth, dorsal surfaces of body lacking dermal spines. Throat black and few dermal spines found on chin and snout (Figure 5B). Cloaca lacks para-cloacal tubercles or glands.
Dorsum dark brown with a few small cream spots and blotches; dorsal surface of limbs dark brown with cream blotches and small spots, particularly on the proximal forelimb; palm of hands marbled brown and pale cream, foot dark brown; belly surface cream, dorsolateral and ventral surfaces with a marbled pale brown and cream pattern; throat dark brown to black. Ventral surface of thighs light brown with a finely reticulated dark pattern over a cream background cream with a few cream spots more evident close to the edges; ventral surfaces of tibia and tarsus finely marbled in light brown with cream, lighter than the dorsal surface. Absence of distinct lines on the body and limbs.
(in mm). SVL 15.7; HDL 3.4; HDL4 2.3; HL 2.7; HW 4.5; ED 1.3; IOD 2.8; IND 1.1; END 1.2; THL 6.5; TBL 6.4; FL 6.9, FAL 3.2; 3FD 0.3; 4TD 0.4.
Measurements data of the type series are given in Table 2 and information of the comparative material are provided in Appendix 1. Overall, the type series agrees with the holotype coloration; one specimen has a mid-dorsal line and a line on the posterior surface of the thighs and also more dermal spines (MZUSP147475). The incomplete occipital fold varied from indistinct to weakly visible laterally (= incomplete). The combined mean thigh and tibia length represents approximately 81% of mean snout-vent length in males, and 77.7% in females; foot length approximately 41.6% of snout-vent length in males and 39.7% in females.
Morphometric measurements (mm) of the type series of Chiasmocleis quilombola sp. n.
| Specimen | Type | Sex | SVL | HL | HW | ED | IOD | IND | END | THL | TBL | FL | 3FD | 4TD | FAL | HDL | HDL4 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| MZUSP147471 | Paratype | Male | 14.4 | 2.8 | 3.8 | 1.0 | 2.6 | 1.1 | 1.5 | 6.1 | 6.3 | 6.2 | 0.3 | 0.3 | 3.0 | 2.8 | 1.8 |
| MZUSP147472 | Paratype | Male | 15.3 | 2.8 | 4.1 | 1.3 | 2.7 | 1.2 | 1.3 | 7.1 | 6.6 | 6.6 | 0.3 | 0.4 | 3.2 | 3.4 | 2.2 |
| MZUSP147473 | Paratype | Male | 16.1 | 3.2 | 3.9 | 1.1 | 2.2 | 1.1 | 1.3 | 6.0 | 6.0 | 6.4 | 0.2 | 0.2 | 3.2 | 3.3 | 2.1 |
| MZUSP147475 | Paratype | Male | 14.5 | 2.8 | 4.6 | 1.3 | 2.6 | 1.3 | 1.3 | 6.6 | 6.4 | 6.7 | 0.4 | 0.4 | 3.3 | 3.5 | 2.2 |
| MZUSP147476 | Paratype | Male | 16.1 | 2.7 | 4.4 | 1.2 | 2.7 | 1.3 | 1.4 | 6.7 | 6.3 | 6.1 | 0.3 | 0.5 | 3.0 | 3.3 | 1.8 |
| MZUSP147478 | Holotype | Male | 15.7 | 2.8 | 4.5 | 1.3 | 2.8 | 1.1 | 1.3 | 6.6 | 6.5 | 6.9 | 0.4 | 0.4 | 3.2 | 3.5 | 2.3 |
| MZUSP147494 | Paratype | Male | 16.6 | 3.1 | 4.6 | 1.4 | 2.6 | 1.2 | 1.4 | 7.0 | 6.9 | 7.6 | 0.3 | 0.4 | 3.3 | 3.8 | 2.4 |
| MZUSP147479 | Paratype | Female | 17.2 | 2.9 | 4.6 | 1.3 | 2.9 | 1.3 | 1.4 | 6.7 | 6.6 | 6.8 | 0.3 | 0.4 | 3.3 | 3.5 | 2.3 |
Abbreviations: SVL = snout-vent length; HDL = hand length; HDL4 = hand length from the base of the thenar tubercle to the tip of the fourth finger; HL = head length; HW = head width; ED = eye diameter; IOD = inter-orbital distance; IND = inter-nostril distance; END = eye-nostril distance; THL = thigh length; TBL = tibia length; FAL = forearm length; FL = foot length; 3FD = diameter of third finger disk; 4TD = diameter of fourth toe disk. For tissues numbers see Appendix 2.
The specific epithet quilombola refers to people who inhabit quilombo communities. Historically, quilombos were communities constituted by and used as refuges for escaped slaves between 1530 and 1815 during colonial Portuguese rule in Brazil. Nowadays in the north of Espírito Santo Estate quilombola communities still remain and maintain alive their traditions, such as quilombola food and craftwork. This species' name is indeclinable.
Chiasmocleis quilombola sp. n. is known from localities between the Doce River and the Mucuri River, e.g., Floresta Nacional do Rio Preto and Parque Estadual de Itaúnas, Municipality of Conceição da Barra; Reserva Biológica Córrego Veado, Municipality of Pinheiros; Reserva Natural, Reserva Biológica de Sooretama, and Cocoa plantations in Povoação, Municipality of Linhares (Appendix 2). The populations assigned to Chiasmocleis lacrimae and Chiasmocleis capixaba at northernmost of Espírito Santo State are allocated to the new taxon Chiasmocleis quilombola sp. n. (Figure 1).
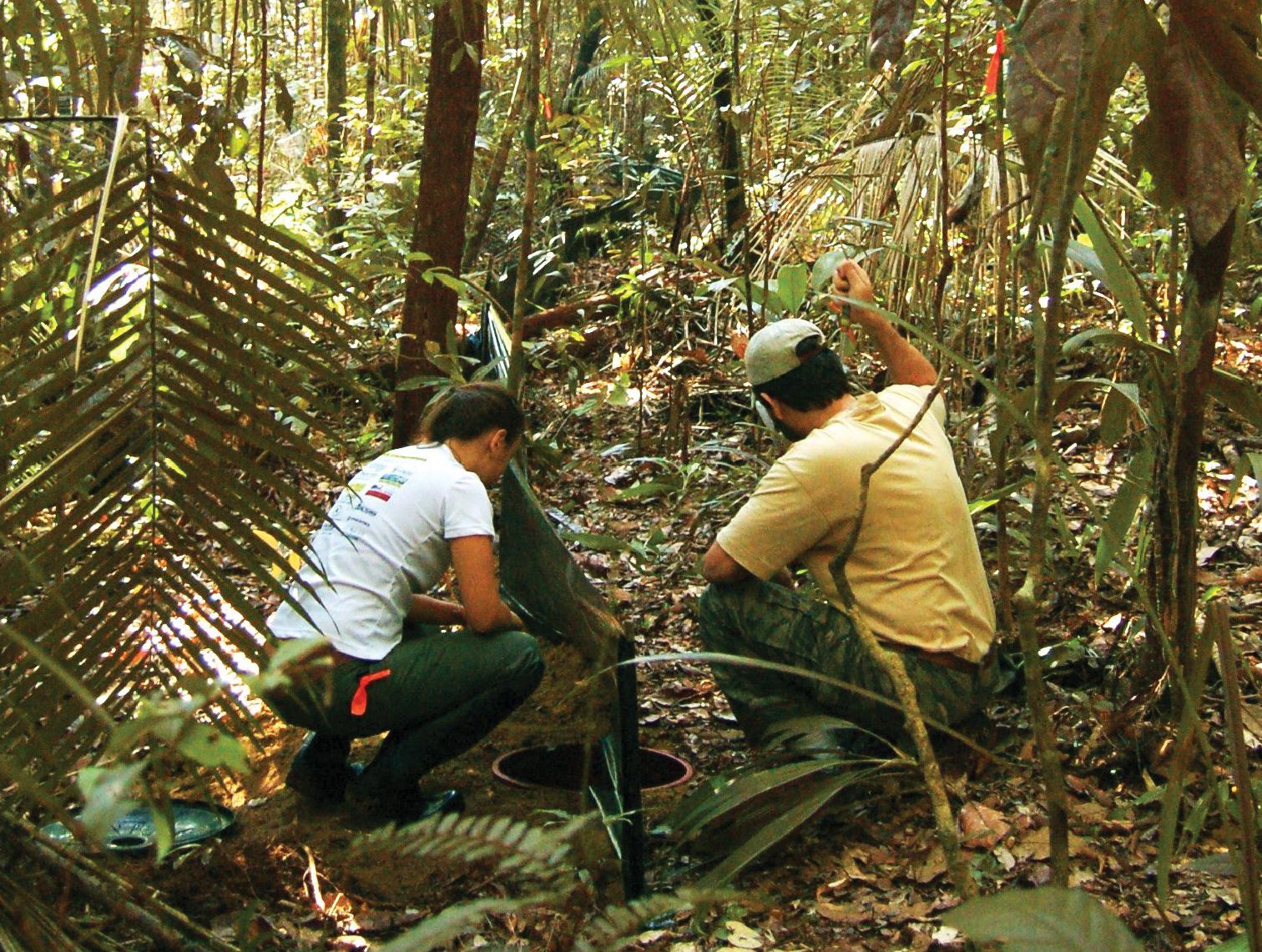
Chiasmocleis quilombola sp. n. was collected in pitfall traps after heavy rains at Floresta Nacional do Rio Preto (Figure 6). The lines of pitfalls were installed at the vicinity of a permanent lagoon and a temporary swamp. The Floresta Nacional do Rio Preto has 2, 830 ha and an elevation between five to 50 m above the sea level. The soil is typical of coastal areas, mostly sand. The area consists of secondary forested areas and plantations with few remnants of primary Atlantic Forest.
Larissa Gaigher and Dr. Yuri Leite inspecting pitfall traps installed at the type locality of Chiasmocleis quilombola sp. n., Floresta Nacional do Rio Preto, Municipality of Conceição da Barra, Espírito Santo State, Brazil.
Chiasmocleis quilombola sp. n. has been misidentified as Chiasmocleis lacrimae and Chiasmocleis capixaba due to an overlap in feet webbing and body size (e.g.,
Genetic distance (p-uncorrected) in the BDNF (upper-right) and in the ND2 (lower-left) among Chiasmocleis quilombola sp. n. and sister species. Values at the diagonal correspond to the genetic distance within species in the ND2.
| Species | Chiasmocleis capixaba | Chiasmocleis lacrimae | Chiasmocleis cordeiroi | Chiasmocleis crucis | Chiasmocleis quilombola sp. n. | Chiasmocleis schubarti | Chiasmocleis sp. |
|---|---|---|---|---|---|---|---|
| Chiasmocleis capixaba | 0.015 | 0.001 | 0.008 | 0.008 | 0.001 | 0.007 | 0.003 |
| Chiasmocleis lacrimae | 0.064 | 0.044 | 0.008 | 0.008 | 0.001 | 0.007 | 0.003 |
| Chiasmocleis cordeiroi | 0.211 | 0.227 | 0.013 | 0.005 | 0.008 | 0.004 | 0.007 |
| Chiasmocleis crucis | 0.182 | 0.198 | 0.108 | 0.015 | 0.008 | 0.005 | 0.007 |
| Chiasmocleis quilombola sp. n. | 0.071 | 0.082 | 0.217 | 0.209 | 0.006 | 0.007 | 0.004 |
| Chiasmocleis schubarti | 0.206 | 0.204 | 0.107 | 0.103 | 0.22 | 0.017 | 0.007 |
| Chiasmocleis sp. | 0.098 | 0.104 | 0.253 | 0.237 | 0.083 | 0.236 | 0.023 |
Chiasmocleis quilombola sp. n. is distinct from Chiasmocleis schubarti (species with which occurs in sympatry) in having smaller snout-vent length, feet slightly webbed, cream ventral surface, and marbled light brown and cream dorsolateral pattern instead of larger snout-vent length, absence of feet webbing and belly pattern roughly marbled in dark brown and light cream in Chiasmocleis schubarti (
Differences between Chiasmocleis capixaba, Chiasmocleis lacrimae, Chiasmocleis quilombola sp. n., and Chiasmocleis sp.
| Species | Chiasmocleis capixaba | Chiasmocleis lacrimae | Chiasmocleis quilombola sp. n. | Chiasmocleis sp. |
|---|---|---|---|---|
| Chiasmocleis capixaba | SVL=15.1 (SD 0.6) HL=2.8 (SD 0.1) THL=6.2 (SD 0.3) TBL=6.1 (SD 0.3) |
Feet webbing | head length; thickness of limbs, fingers, and toes; dermal spines | Feet webbing; thickness of limbs; dermal spines |
| Chiasmocleis lacrimae | mtDNA (ND2: 6.4%, 16S: 1.3%, 12S: 1.8%) nuDNA (BNDF: haplotype sharing) | SVL=16.1 (SD 0.9) HL=3.3 (SD 0.2) THL=6.6 (SD 0.4) TBL=6.5 (SD 0.3) |
head and limb length; feet webbing; thickness of limbs; dermal spines | body size; thickness; dermal spines |
| Chiasmocleis quilombola sp. n. | mtDNA (ND2: 7.1%, 16S: 0.8%, 12S: 1.1%) nuDNA (BNDF: haplotype sharing) | mtDNA (ND2: 8.2%, 16S: 0.8%, 12S: 1.8%) nuDNA (BNDF: haplotype sharing) | SVL=14 (SD 1.4) HL=2.6 (SD 0.2) THL=5.8 (SD 0.7) TBL=5.6 (SD 0.6) |
Feet webbing; thickness of limbs |
| Chiasmocleis sp. | mtDNA (ND2: 9.8%, 16S: 2.2%, 12S: 1.2%) nuDNA (BNDF: no haplotype sharing) | mtDNA (ND2: 10%, 16S: 2.3%, 12S: 2.1%) nuDNA (BNDF: no haplotype sharing) | mtDNA (ND2: 8.3%, 16S: 1.6%, 12S: 2.1%) nuDNA (BNDF: haplotype sharing) | SVL=15.3 (SD 0.7) HL=3.4 (SD 0.1) THL=6 (SD 0.3) TBL=6.2 (SD 0.3) |
SVL snout-vent length; HL head length; THL tight length; TBL tibia length; SD standard deviation.
Chiasmocleis quilombola sp. n. are distinguished from other Chiasmocleis species by: 1) four externally evident fingers and five toes distinguishes it from Chiasmocleis antenori, Chiasmocleis carvalhoi, and Chiasmocleis tridactyla (digit reduction;
Chiasmocleis quilombola sp. n. occurs in sympatry with Chiasmocleis schubarti at the Floresta Nacional do Rio Preto, Municipality of Conceição da Barra, and at the Reserva Biológica Córrego Veado, Municipality of Pinheiros; it also occurs with Chiasmocleis capixaba and Chiasmocleis schubarti at the Reserva Natural Vale, Reserva Biológica de Sooretama, and at Cocoa plantations in Povoação, sites in the Municipality of Linhares. The new species is allopatric to Chiasmocleis sp. and Chiasmocleis lacrimae (Figure 1). We did not have access to tissues samples of Chiasmocleis capixaba from Nova Viçosa, Bahia State (
Cryptic species have challenged our ability to assess current levels of biodiversity. Anuran taxonomy has used various data sources to describe the species diversity, e.g., advertisement calls, external morphology, osteology, tadpoles, ecology, molecular data, karyotypes (
The new species occupy coastal areas North of Espírito Santo State, a region that is under strong human pressure. Therefore, marine and coastal communities are susceptible to impacts of proposed modifications in the landscape for the exploitation of mineral resources. In this context, Chiasmocleis quilombola sp. n. may face imminent threat of habitat loss, as consequence of the deforestation and intensive occupation of the space by human activities.
We are thankful to L. Costa, Y. Leite, and the Laboratório de Mastozoologia e Biogeografia team, for field assistance and tissues samples. L. Chagas and staffs of the Floresta Nacional do Rio Preto for valuable help during field surveys. M. T. Rodrigues, R. C. Amaro, C. F. B. Haddad, P. Rocha, M. Napoli, H. Zaher, M. Solé, V. Fagundes, J. L. Gasparini, J. P. Pombal Jr., and P. Passos for providing tissue samples and permission to examine specimens under their care. D. Baêta and A. C. Calijorne for hosting JFRT during visit to the Museu Nacional do Rio de Janeiro. R. Ferreira, J. L. Gasparini, M. Vences, R. A. Pyron, F. Andreone, and two anonymous referees for suggestions on the manuscript. JFRT acknowledge support by the Science without Borders program (CAPES/Brazil), The George Washington University, and award NSF-DEB 1144692 to R. O. de Sá.
Examined material and measurements from males included in the morphological analyses.
| Chiasmocleis | Numbers | Locality | State | SVL | HL | HW | ED | IOD | IND | END | THL | TBL | FL | 3FD | 4TD | FAL | HDL | HDL4 |
| Chiasmocleis capixaba | MNRJ17514 | Aracruz | ES | 14.77 | 2.91 | 4.04 | 1.18 | 2.57 | 1.16 | 1.21 | 6.19 | 6.34 | 6.43 | 0.35 | 0.44 | 2.98 | 3.36 | 2.08 |
| Chiasmocleis capixaba | MNRJ17515 | Aracruz | ES | 13.98 | 2.84 | 4.44 | 1.28 | 2.71 | 1.21 | 1.39 | 6.17 | 6.13 | 6.58 | 0.29 | 0.46 | 3.07 | 3.28 | 1.94 |
| Chiasmocleis capixaba | MNRJ17516 | Aracruz | ES | 14.47 | 2.93 | 3.98 | 1.18 | 2.64 | 1.19 | 1.32 | 6.08 | 5.96 | 6.64 | 0.30 | 0.42 | 3.10 | 3.36 | 2.16 |
| Chiasmocleis capixaba | MNRJ17517 | Aracruz | ES | 14.70 | 3.00 | 4.10 | 1.36 | 2.61 | 1.29 | 1.26 | 6.23 | 6.10 | 6.74 | 0.33 | 0.51 | 3.15 | 3.55 | 2.32 |
| Chiasmocleis capixaba | MNRJ17518 | Aracruz | ES | 16.02 | 3.13 | 4.39 | 1.32 | 2.64 | 1.21 | 1.45 | 6.61 | 6.80 | 7.45 | 0.38 | 0.48 | 3.62 | 3.86 | 2.33 |
| Chiasmocleis capixaba | MNRJ17519 | Aracruz | ES | 15.40 | 2.99 | 3.95 | 1.36 | 2.63 | 1.23 | 1.56 | 6.05 | 6.31 | 6.28 | 0.33 | 0.45 | 3.38 | 3.17 | 2.09 |
| Chiasmocleis capixaba | MNRJ17520 | Aracruz | ES | 14.94 | 2.76 | 3.82 | 1.26 | 2.66 | 1.20 | 1.36 | 5.71 | 6.06 | 5.90 | 0.41 | 0.47 | 2.96 | 3.15 | 1.80 |
| Chiasmocleis capixaba | MNRJ17535 | Aracruz | ES | 15.23 | 2.96 | 3.83 | 0.97 | 2.39 | 1.18 | 1.07 | 6.18 | 5.76 | 5.97 | 0.38 | 0.42 | 2.76 | 3.05 | 2.13 |
| Chiasmocleis capixaba | MNRJ17536 | Aracruz | ES | 15.24 | 2.50 | 3.92 | 1.23 | 2.60 | 1.16 | 1.41 | 5.83 | 5.79 | 6.63 | 0.33 | 0.45 | 2.93 | 3.52 | 2.19 |
| Chiasmocleis capixaba | MNRJ17895 | Aracruz | ES | 15.46 | 2.89 | 4.32 | 1.12 | 2.56 | 1.22 | 1.21 | 6.62 | 6.30 | 6.46 | 0.33 | 0.47 | 3.25 | 3.34 | 2.15 |
| Chiasmocleis capixaba | MNRJ22962 | Reserva Natural Vale | ES | 14.83 | 3.02 | 4.44 | 1.26 | 2.67 | 1.18 | 1.38 | 6.29 | 6.08 | 6.37 | 0.36 | 0.44 | 3.12 | 3.34 | 2.08 |
| Chiasmocleis capixaba | MNRJ22966 | Reserva Natural Vale | ES | 15.05 | 2.99 | 4.41 | 1.37 | 2.71 | 1.19 | 1.43 | 6.71 | 6.20 | 6.25 | 0.36 | 0.52 | 3.18 | 3.15 | 1.72 |
| Chiasmocleis capixaba | MZUSP147468 | ReBio Duas Bocas | ES | 15.88 | 3.05 | 4.40 | 1.27 | 2.87 | 1.25 | 1.41 | 6.84 | 6.53 | 6.92 | 0.43 | 0.43 | 3.06 | 3.44 | 2.14 |
| Chiasmocleis capixaba | MZUSP147469 | ReBio Duas Bocas | ES | 16.23 | 3.02 | 4.61 | 1.05 | 2.99 | 1.31 | 1.68 | 6.80 | 6.71 | 7.05 | 0.48 | 0.48 | 3.28 | 3.29 | 2.30 |
| Chiasmocleis lacrimae | MNRJ17480 | Horto Florestal | RJ | 16.20 | 3.40 | 4.40 | 1.10 | 2.70 | 0.90 | 1.40 | 6.50 | 6.50 | 7.30 | 0.30 | 0.40 | 3.30 | 3.60 | 2.50 |
| Chiasmocleis lacrimae | MNRJ17481 | Horto Florestal | RJ | 15.60 | 3.30 | 4.10 | 1.10 | 2.60 | 1.00 | 1.40 | 6.20 | 6.30 | 7.20 | 0.40 | 0.40 | 3.50 | 3.60 | 2.60 |
| Chiasmocleis lacrimae | MNRJ17482 | Horto Florestal | RJ | 16.20 | 3.50 | 2.60 | 1.30 | 2.60 | 1.00 | 1.90 | 6.50 | 6.50 | 7.10 | 0.40 | 0.50 | 3.30 | 3.80 | 2.50 |
| Chiasmocleis lacrimae | MNRJ17484 | Horto Florestal | RJ | 19.15 | 3.57 | 4.82 | 1.35 | 2.84 | 1.44 | 1.69 | 7.45 | 7.35 | 8.23 | 0.34 | 0.36 | 3.56 | 4.23 | 2.69 |
| Chiasmocleis lacrimae | MNRJ17485 | Horto Florestal | RJ | 16.10 | 3.70 | 4.30 | 1.20 | 2.50 | 0.90 | 1.60 | 6.90 | 6.70 | 7.50 | 0.30 | 0.40 | 3.40 | 3.60 | 2.40 |
| Chiasmocleis lacrimae | MNRJ17486 | Horto Florestal | RJ | 15.07 | 3.00 | 4.33 | 1.22 | 2.52 | 1.22 | 1.50 | 6.30 | 6.22 | 6.68 | 0.30 | 0.43 | 3.19 | 3.44 | 2.24 |
| Chiasmocleis lacrimae | MNRJ17487 | Horto Florestal | RJ | 16.10 | 3.33 | 4.38 | 1.19 | 2.91 | 1.30 | 1.68 | 6.94 | 6.78 | 7.16 | 0.27 | 0.39 | 3.46 | 3.53 | 2.08 |
| Chiasmocleis lacrimae | MNRJ17488 | Horto Florestal | RJ | 15.90 | 3.20 | 3.80 | 1.10 | 2.50 | 0.90 | 1.30 | 5.80 | 6.20 | 6.60 | 0.30 | 0.40 | 3.00 | 3.30 | 2.20 |
| Chiasmocleis lacrimae | MNRJ17489 | Horto Florestal | RJ | 15.40 | 3.10 | 3.90 | 1.20 | 2.20 | 0.90 | 1.30 | 6.10 | 6.10 | 6.90 | 0.30 | 0.40 | 3.10 | 3.40 | 2.10 |
| Chiasmocleis lacrimae | MNRJ17490 | Horto Florestal | RJ | 16.10 | 3.30 | 4.00 | 1.10 | 2.50 | 0.90 | 1.40 | 6.70 | 7.10 | 7.70 | 0.40 | 0.40 | 3.50 | 4.20 | 2.70 |
| Chiasmocleis lacrimae | MNRJ17491 | Horto Florestal | RJ | 15.26 | 2.90 | 4.75 | 1.21 | 2.92 | 1.47 | 1.49 | 6.59 | 6.46 | 7.13 | 0.32 | 0.41 | 3.34 | 3.78 | 2.41 |
| Chiasmocleis lacrimae | MNRJ17492 | Horto Florestal | RJ | 15.50 | 3.90 | 4.60 | 1.30 | 2.50 | 1.20 | 1.50 | 6.10 | 6.30 | 6.50 | 0.40 | 0.40 | 3.10 | 3.70 | 2.50 |
| Chiasmocleis lacrimae | MNRJ17498 | Horto Florestal | RJ | 17.10 | 3.70 | 4.70 | 1.20 | 2.50 | 1.10 | 1.40 | 6.70 | 6.80 | 6.90 | 0.40 | 0.50 | 3.40 | 3.40 | 2.50 |
| Chiasmocleis lacrimae | MNRJ17505 | Horto Florestal | RJ | 16.80 | 3.42 | 4.62 | 1.41 | 2.84 | 1.32 | 1.71 | 7.19 | 6.88 | 7.43 | 0.33 | 0.38 | 3.50 | 3.87 | 2.42 |
| Chiasmocleis lacrimae | MNRJ17506 | Horto Florestal | RJ | 16.95 | 3.30 | 4.84 | 1.36 | 3.00 | 1.35 | 1.62 | 7.19 | 6.80 | 7.32 | 0.41 | 0.43 | 3.60 | 4.09 | 2.63 |
| Chiasmocleis lacrimae | MNRJ17507 | Horto Florestal | RJ | 15.16 | 3.14 | 4.43 | 1.29 | 2.71 | 1.24 | 1.45 | 6.06 | 6.13 | 6.28 | 0.34 | 0.41 | 3.12 | 3.51 | 2.22 |
| Chiasmocleis lacrimae | MNRJ17565 | Horto Florestal | RJ | 16.66 | 2.97 | 4.42 | 1.36 | 2.79 | 1.30 | 1.47 | 7.05 | 6.69 | 7.14 | 0.49 | 0.43 | 3.23 | 3.97 | 2.63 |
| Chiasmocleis lacrimae | MNRJ66497 | Mimoso do Sul | ES | 16.60 | 3.14 | 4.77 | 1.40 | 2.89 | 1.31 | 1.48 | 6.69 | 6.49 | 6.90 | 0.39 | 0.43 | 3.55 | 3.80 | 2.32 |
| Chiasmocleis quilombola sp. n. | MNRJ29057 | Povoação | ES | 12.72 | 2.70 | 3.89 | 1.05 | 2.42 | 1.08 | 1.20 | 5.35 | 5.05 | 5.24 | 0.30 | 0.39 | 2.65 | 2.79 | 1.79 |
| Chiasmocleis quilombola sp. n. | MNRJ29058 | Povoação | ES | 13.23 | 2.68 | 3.90 | 1.12 | 2.47 | 1.10 | 1.12 | 5.28 | 4.91 | 5.17 | 0.34 | 0.42 | 2.52 | 2.78 | 1.73 |
| Chiasmocleis quilombola sp. n. | MNRJ29059 | Povoação | ES | 13.56 | 2.53 | 3.63 | 1.15 | 2.37 | 1.08 | 1.12 | 5.36 | 5.25 | 5.33 | 0.29 | 0.39 | 2.80 | 3.00 | 1.89 |
| Chiasmocleis quilombola sp. n. | MNRJ29060 | Povoação | ES | 12.59 | 2.55 | 3.79 | 1.08 | 2.41 | 1.08 | 1.16 | 5.12 | 4.67 | 4.56 | 0.32 | 0.32 | 2.48 | 2.48 | 1.67 |
| Chiasmocleis quilombola sp. n. | MNRJ29073 | Povoação | ES | 13.76 | 2.53 | 3.94 | 1.25 | 2.43 | 1.18 | 1.29 | 5.67 | 5.63 | 5.88 | 0.41 | 0.41 | 2.88 | 3.07 | 1.85 |
| Chiasmocleis quilombola sp. n. | MNRJ29074 | Povoação | ES | 13.47 | 2.33 | 3.72 | 1.18 | 2.27 | 1.08 | 1.15 | 5.24 | 5.27 | 5.60 | 0.32 | 0.40 | 2.73 | 3.02 | 1.81 |
| Chiasmocleis quilombola sp. n. | MBML2858 | Povoação | ES | 13.47 | 2.54 | 3.94 | 0.88 | 2.04 | 0.82 | 1.03 | 5.11 | 5.39 | 5.50 | 0.33 | 0.43 | 2.61 | 2.91 | 1.72 |
| Chiasmocleis quilombola sp. n. | MBML2866 | Povoação | ES | 12.15 | 2.24 | 3.34 | 1.05 | 2.16 | 0.72 | 0.95 | 5.26 | 5.11 | 5.33 | 0.27 | 0.39 | 2.55 | 2.75 | 1.62 |
| Chiasmocleis quilombola sp. n. | MBML2863 | Povoação | ES | 12.30 | 2.34 | 3.75 | 0.82 | 2.00 | 0.83 | 0.93 | 5.22 | 5.32 | 5.42 | 0.31 | 0.48 | 2.50 | 2.63 | 1.59 |
| Chiasmocleis quilombola sp. n. | MZUSP147473 | Flona Rio Preto | ES | 16.07 | 3.18 | 3.87 | 1.09 | 2.18 | 1.09 | 1.33 | 6.01 | 6.00 | 6.41 | 0.24 | 0.23 | 3.15 | 3.30 | 2.07 |
| Chiasmocleis quilombola sp. n. | MZUSP147478 | Flona Rio Preto | ES | 15.71 | 2.75 | 4.52 | 1.30 | 2.80 | 1.13 | 1.25 | 6.55 | 6.47 | 6.91 | 0.35 | 0.44 | 3.21 | 3.45 | 2.33 |
| Chiasmocleis quilombola sp. n. | MZUSP147471 | Flona Rio Preto | ES | 14.43 | 2.75 | 3.83 | 1.03 | 2.57 | 1.11 | 1.51 | 6.07 | 6.27 | 6.23 | 0.25 | 0.26 | 2.99 | 2.84 | 1.83 |
| Chiasmocleis quilombola sp. n. | MZUSP147475 | Flona Rio Preto | ES | 14.48 | 2.79 | 4.55 | 1.28 | 2.55 | 1.28 | 1.25 | 6.63 | 6.41 | 6.73 | 0.36 | 0.36 | 3.26 | 3.48 | 2.16 |
| Chiasmocleis quilombola sp. n. | MZUSP147494 | Flona Rio Preto | ES | 16.55 | 3.14 | 4.63 | 1.36 | 2.55 | 1.24 | 1.36 | 7.02 | 6.89 | 7.59 | 0.25 | 0.37 | 3.33 | 3.80 | 2.40 |
| Chiasmocleis quilombola sp. n. | MZUSP147476 | Flona Rio Preto | ES | 16.09 | 2.70 | 4.36 | 1.24 | 2.65 | 1.31 | 1.36 | 6.73 | 6.26 | 6.11 | 0.32 | 0.47 | 2.95 | 3.33 | 1.83 |
| Chiasmocleis quilombola sp. n. | MZUSP147472 | Flona Rio Preto | ES | 15.28 | 2.80 | 4.08 | 1.33 | 2.67 | 1.24 | 1.33 | 7.08 | 6.55 | 6.60 | 0.34 | 0.41 | 3.20 | 3.40 | 2.22 |
| sp. | MTR13495 | Trancoso | BA | 15.92 | 3.39 | 4.07 | 1.06 | 2.65 | 0.94 | 1.43 | 6.37 | 6.16 | 6.77 | 0.27 | 0.42 | 3.60 | 3.45 | 2.06 |
| sp. | MTR13547 | Trancoso | BA | 13.80 | 3.53 | 4.09 | 0.97 | 2.36 | 1.00 | 1.35 | 5.59 | 5.68 | 5.72 | 0.27 | 0.35 | 2.95 | 3.18 | 2.27 |
| sp. | MTR13545 | Trancoso | BA | 15.07 | 3.51 | 4.24 | 1.10 | 2.43 | 1.03 | 1.35 | 5.91 | 6.41 | 6.78 | 0.29 | 0.38 | 2.95 | 3.40 | 2.35 |
| sp. | MTR13590 | Trancoso | BA | 15.48 | 3.48 | 3.99 | 1.17 | 2.51 | 0.80 | 1.32 | 6.01 | 6.22 | 6.69 | 0.30 | 0.45 | 3.11 | 3.73 | 2.41 |
| sp. | MTR13565 | Trancoso | BA | 16.27 | 3.78 | 4.31 | 0.95 | 2.59 | 1.04 | 1.61 | 5.64 | 6.33 | 6.56 | 0.33 | 0.46 | 3.20 | 3.68 | 2.67 |
| sp. | MTR13548 | Trancoso | BA | 15.36 | 3.34 | 4.57 | 1.06 | 2.52 | 1.12 | 1.09 | 6.44 | 6.62 | 6.72 | 0.30 | 0.40 | 3.19 | 3.64 | 2.36 |
| sp. | MTR13546 | Trancoso | BA | 15.53 | 3.10 | 4.40 | 1.24 | 2.30 | 0.91 | 1.46 | 6.02 | 5.96 | 6.56 | 0.31 | 0.40 | 2.90 | 3.30 | 2.14 |
| sp. | MTR13489 | Trancoso | BA | 15.34 | 3.53 | 4.05 | 1.16 | 2.35 | 1.00 | 1.36 | 6.67 | 6.54 | 6.73 | 0.27 | 0.44 | 3.40 | 3.46 | 2.40 |
Samples included in the molecular analyses and genbank numbers. Numbers in bold were retrieved from previous studies.
| Chiasmocleis | Voucher | Tissue | Locality | State | 12S | 16S | ND2 | BDNF |
| Chiasmocleis capixaba | MZUSP147497 | CTA1861 | ReBio Duas Bocas | ES | KM111721 | KM111817 | JQ410706 | KM111908 |
| Chiasmocleis capixaba | MZUSP147498 | CTA1863 | ReBio Duas Bocas | ES | KM111722 | KM111818 | JQ410707 | KM111909 |
| Chiasmocleis capixaba | MZUSP147499 | CTA1864 | ReBio Duas Bocas | ES | KM111723 | KM111819 | JQ410708 | KM111910 |
| Chiasmocleis capixaba | MZUSP147468 | CTA1865 | ReBio Duas Bocas | ES | KM111724 | KM111820 | KM111992 | KM111911 |
| Chiasmocleis capixaba | MZUSP147482 | CTA1869 | ReBio Duas Bocas | ES | KM111725 | KM111821 | KM111993 | KM111912 |
| Chiasmocleis capixaba | MZUSP147469 | CTA1870 | ReBio Duas Bocas | ES | KM111726 | KM111822 | KM111994 | KM111913 |
| Chiasmocleis capixaba | MZUSP147500 | CTA1871 | ReBio Duas Bocas | ES | KM111727 | KM111823 | JQ410709 | KM111914 |
| Chiasmocleis capixaba | MZUSP147510 | CTA1872 | Serra | ES | KM111728 | KM111824 | JQ410685 | KM111915 |
| Chiasmocleis capixaba | MZUSP147512 | CTA1874 | Serra | ES | KM111729 | KM111825 | JQ410690 | KM111916 |
| Chiasmocleis capixaba | MZUSP147513 | CTA1875 | Serra | ES | KM111730 | KM111826 | JQ410691 | KM111917 |
| Chiasmocleis capixaba | JFT479 | CTA1876 | Serra | ES | KM111731 | KM111827 | JQ410692 | KM111918 |
| Chiasmocleis capixaba | MZUSP147514 | CTA1877 | Serra | ES | KM111732 | KM111828 | JQ410693 | KM111919 |
| Chiasmocleis capixaba | JFT483 | CTA1879 | Serra | ES | KM111733 | KM111829 | JQ410694 | KM111920 |
| Chiasmocleis capixaba | JFT499 | CTA1881 | Serra | ES | KM111734 | KM111830 | JQ410695 | KM111921 |
| Chiasmocleis capixaba | MZUSP147520 | CTA1893 | Serra | ES | KM111735 | KM111831 | JQ410700 | KM111922 |
| Chiasmocleis capixaba | MZUSP147521 | CTA1894 | Serra | ES | KM111736 | KM111832 | JQ410701 | KM111923 |
| Chiasmocleis capixaba | MZUSP147523 | CTA1896 | Serra | ES | KM111737 | KM111833 | JQ410702 | KM111924 |
| Chiasmocleis capixaba | MTR12276 | CTMZ6907 | FloNa dos Goytacazes | ES | KM111738 | KM111834 | - | KM111925 |
| Chiasmocleis capixaba | MTR12296 | CTMZ6908 | FloNa dos Goytacazes | ES | KM111739 | KM111835 | JQ410688 | KM111926 |
| Chiasmocleis capixaba | MTR12407 | CTMZ6912 | Reserva Natural Vale | ES | KM111740 | KM111836 | - | - |
| Chiasmocleis capixaba | MTR12484 | CTMZ6920 | Reserva Natural Vale | ES | KM111741 | KM111837 | - | KM111927 |
| Chiasmocleis capixaba | MTR12485 | CTMZ6921 | Reserva Natural Vale | ES | KM111742 | KM111838 | - | KM111928 |
| Chiasmocleis capixaba | - | CTMZ6939 | Guarapari | ES | KM111743 | KM111839 | - | KM111929 |
| Chiasmocleis capixaba | MTR12076 | MTR12076 | Reserva Natural Vale | ES | KM111744 | KM111840 | JQ410687 | KM111930 |
| Chiasmocleis capixaba | MTR12297 | MTR12297 | FloNa dos Goytacazes | ES | KM111745 | KM111841 | JQ410689 | KM111931 |
| Chiasmocleis cordeiroi | CFBH32057 | CFBH15784 | Ilhéus | BA | KM111759 | KM111852 | KM111995 | KM111939 |
| Chiasmocleis cordeiroi | MZUSP147496 | CTA1935 | Ituberá | BA | KM111760 | KM111853 | KM111996 | KM111940 |
| Chiasmocleis cordeiroi | MTR22122 | MTR22122 | EE Wenceslau Guimarães | BA | KM111761 | KM111854 | KM111997 | KM111941 |
| Chiasmocleis cordeiroi | MTR22123 | MTR22123 | EE Wenceslau Guimarães | BA | KM111762 | KM111855 | KM111998 | KM111942 |
| Chiasmocleis cordeiroi | PEU137 | PEU137 | Jaguaripe | BA | KM111763 | KM111856 | KM111999 | KM111943 |
| Chiasmocleis cordeiroi | PEU146 | PEU146 | Jaguaripe | BA | KM111764 | KM111857 | KM112000 | KM111944 |
| Chiasmocleis crucis | MTR6001 | CTMZ6898 | Serra do Teimoso | BA | KM111765 | KM111858 | KM112001 | KM111945 |
| Chiasmocleis crucis | - | CTMZ6900 | Ilhéus | BA | KM111766 | KM111859 | KM112002 | KM111946 |
| Chiasmocleis crucis | - | CTMZ6901 | Ilhéus | BA | KM111767 | KM111860 | KM112003 | KM111947 |
| Chiasmocleis crucis | MTR16070 | MTR16070 | Serra Bonita | BA | KM111768 | KM111861 | KM112004 | KM111948 |
| Chiasmocleis lacrimae | - | CFBH73 | Picinguaba | SP | KM111748 | KC180040 | JQ410715 | KC180202 |
| Chiasmocleis lacrimae | CFBH17495 | CFBH7361 | Ilha de São Sebastião | SP | KM111749 | KM111844 | - | |
| Chiasmocleis lacrimae | - | CFBH76 | Picinguaba | SP | KM111750 | KC180063 | JQ410714 | KC180163 |
| Chiasmocleis lacrimae | JFT981 | CTA1934 | Matada Usina Paineiras | ES | KM111751 | KM111845 | JQ410710 | KM111932 |
| Chiasmocleis lacrimae | - | CTMZ6946 | Bertioga | SP | KM111752 | KM111846 | - | KM111933 |
| Chiasmocleis lacrimae | RN7003 | CTRN173 | Angra dos Reis | RJ | KM111753 | KM111847 | - | - |
| Chiasmocleis lacrimae | RN7004 | CTRN174 | Angra dos Reis | RJ | KM111754 | KM111848 | - | KM111934 |
| Chiasmocleis lacrimae | RN7005 | CTRN175 | Angra dos Reis | RJ | KM111755 | KM111849 | - | KM111935 |
| Chiasmocleis lacrimae | MNRJ47477 | MNRJ47477 | ReBio União | RJ | KM111746 | KM111842 | - | - |
| Chiasmocleis lacrimae | MNRJ48415 | MNRJ48415 | ReBio União | RJ | KM111747 | KM111843 | - | - |
| Chiasmocleis lacrimae | MNRJ49302 | MNRJ49302 | Cachoeiras de Macacu | ES | KM111756 | KM111850 | JQ410712 | KM111936 |
| Chiasmocleis lacrimae | MNRJ60744 | MNRJ60744 | Duque de Caxias | RJ | KM111757 | - | JQ410713 | KM111937 |
| Chiasmocleis lacrimae | MNRJ66494 | MNRJ66494 | Mimoso do Sul | ES | KM111758 | KM111851 | JQ410711 | KM111938 |
| Chiasmocleis leucosticta | CFBH19029 | CFBH8594 | PE Ilha do Cardoso | SP | KM111769 | KM111862 | - | - |
| Chiasmocleis leucosticta | MZUSP136053 | CTMZ2485 | PE Carlos Botelho | SP | KM111770 | KM111863 | - | - |
| Chiasmocleis leucosticta | MZUSP136055 | CTMZ2493 | PE Carlos Botelho | SP | KM111771 | - | - | KM111949 |
| Chiasmocleis leucosticta | MZUSP136059 | CTMZ2497 | PE Carlos Botelho | SP | - | KM111864 | - | KM111950 |
| Chiasmocleis leucosticta | MTR7128 | CTMZ6943 | Fazenda Intervales | SP | - | - | KM112005 | KM111951 |
| Chiasmocleis leucosticta | - | CTMZ6944 | Piedade | SP | KM111772 | - | KM112006 | - |
| Chiasmocleis mantiqueira | - | CTMZ6891 | Serra do Brigadeiro | MG | KM111773 | KM111865 | - | KM111952 |
| Chiasmocleis mantiqueira | UFMG-A9643 | UFMG-T1802 | Ouro Branco | MG | KM111774 | KM111866 | KM112007 | - |
| Chiasmocleis mantiqueira | UFMG-A9659 | UFMG-T1804 | Ouro Branco | MG | KM111775 | KM111867 | - | KM111953 |
| Chiasmocleis mantiqueira | UFMG-A9651 | UFMG-T1810 | Ouro Branco | MG | KM111776 | KM111868 | - | KM111954 |
| Chiasmocleis mantiqueira | UFMG-A9656 | UFMG-T1815 | Ouro Branco | MG | KM111777 | KM111869 | - | KM111955 |
| Chiasmocleis quilombola sp. n. | - | CFBH1437 | ReBio Sooretama | ES | KM111778 | KC180044 | - | KC180193 |
| Chiasmocleis quilombola sp. n. | - | CFBH1438 | ReBio Sooretama | ES | KM111779 | KC179977 | - | KC180168 |
| Chiasmocleis quilombola sp. n. | CFBH19471 | CFBH9055 | Povoação | ES | KM111780 | KM111870 | KM112008 | KM111956 |
| Chiasmocleis quilombola sp. n. | CFBH18076 | CFBH9082 | Povoação | ES | KM111781 | KM111871 | KM112009 | KM111957 |
| Chiasmocleis quilombola sp. n. | CFBH18077 | CFBH9083 | Povoação | ES | KM111782 | KM111872 | KM112010 | KM111958 |
| Chiasmocleis quilombola sp. n. | JFT831 | CTA1906 | FloNa do Rio Preto | ES | KM111783 | KM111873 | JQ410669 | KM111959 |
| Chiasmocleis quilombola sp. n. | MZUSP147471 | CTA1907 | FloNa do Rio Preto | ES | KM111784 | KM111874 | JQ410670 | KM111960 |
| Chiasmocleis quilombola sp. n. | MZUSP147472 | CTA1908 | FloNa do Rio Preto | ES | KM111785 | KM111875 | JQ410671 | KM111961 |
| Chiasmocleis quilombola sp. n. | MZUSP147473 | CTA1909 | FloNa do Rio Preto | ES | KM111786 | KM111876 | JQ410672 | KM111962 |
| Chiasmocleis quilombola sp. n. | MZUSP147474 | CTA1918 | FloNa do Rio Preto | ES | KM111787 | KM111877 | JQ410673 | KM111963 |
| Chiasmocleis quilombola sp. n. | MZUSP147475 | CTA1919 | FloNa do Rio Preto | ES | KM111788 | KM111878 | JQ410674 | KM111964 |
| Chiasmocleis quilombola sp. n. | MZUSP147494 | CTA1923 | FloNa do Rio Preto | ES | KM111789 | KM111879 | JQ410677 | KM111965 |
| Chiasmocleis quilombola sp. n. | MZUSP147479 | CTA1929 | FloNa do Rio Preto | ES | KM111790 | KM111880 | JQ410679 | KM111966 |
| Chiasmocleis quilombola sp. n. | MZUSP147480 | CTA1931 | FloNa do Rio Preto | ES | KM111791 | KM111881 | JQ410680 | KM111967 |
| Chiasmocleis quilombola sp. n. | MZUSP147493 | CTA1933 | FloNa do Rio Preto | ES | KM111792 | KM111882 | JQ410681 | KM111968 |
| Chiasmocleis quilombola sp. n. | JFT990 | CTA1938 | PE de Itaúnas | ES | - | KM111883 | - | KM111969 |
| Chiasmocleis quilombola sp. n. | MTR12017 | CTMZ6903 | Reserva Natural Vale | ES | KM111793 | KM111884 | - | KM111970 |
| Chiasmocleis quilombola sp. n. | MTR12077 | CTMZ6905 | Reserva Natural Vale | ES | KM111794 | KM111885 | - | - |
| Chiasmocleis quilombola sp. n. | MTR12470 | CTMZ6916 | Reserva Natural Vale | ES | KM111795 | KM111886 | - | KM111971 |
| Chiasmocleis quilombola sp. n. | MTR12471 | CTMZ6917 | Reserva Natural Vale | ES | KM111796 | KM111887 | - | KM111972 |
| Chiasmocleis quilombola sp. n. | MTR21527 | LGA3267 | ReBio Córrego Veado | ES | KM111797 | KM111888 | JQ410668 | KM111973 |
| Chiasmocleis schubarti | CFBH9331 | CFBH2078 | ReBio Sooretama | ES | KM111798 | KC180071 | KM112011 | KC180122 |
| Chiasmocleis schubarti | CFBH18075 | CFBH9060 | Povoação | ES | KM111799 | KM111889 | KM112012 | KM111974 |
| Chiasmocleis schubarti | CFBH 22501 | CTA1860 | ReBio Duas Bocas | ES | KM111800 | KM111890 | KM112013 | KM111975 |
| Chiasmocleis schubarti | MZUSP147485 | CTA1887 | FloNa do Rio Preto | ES | KM111801 | KM111891 | KM112014 | KM111976 |
| Chiasmocleis schubarti | MZUSP147487 | CTA1924 | FloNa do Rio Preto | ES | KM111802 | KM111892 | KM112015 | KM111977 |
| Chiasmocleis schubarti | MTR12094 | CTMZ6906 | FloNa dos Goytacazes | ES | KM111803 | KM111893 | KM112016 | KM111978 |
| Chiasmocleis schubarti | LGA2630 | LGA2630 | ReBio Córrego Veado | ES | KM111804 | KM111894 | JQ410661 | KM111979 |
| Chiasmocleis schubarti | MTR12266 | MTR12266 | FloNa dos Goytacazes | ES | KM111805 | KM111895 | KM112017 | KM111980 |
| Chiasmocleis schubarti | MTR17524 | MTR17524 | PE do Rio Doce | MG | KM111806 | KM111896 | KM112018 | KM111981 |
| Chiasmocleis schubarti | MTR17571 | MTR17571 | PE do Rio Doce | MG | KM111807 | KM111897 | KM112019 | KM111982 |
| sp. | - | CFBH15818 | Porto Seguro | BA | - | KM111898 | - | KM111983 |
| sp. | MTR13466 | CTMZ6923 | Trancoso | BA | KM111808 | KM111899 | JQ410665 | KM111984 |
| sp. | MTR13489 | CTMZ6924 | Trancoso | BA | KM111809 | KM111900 | JQ410666 | KM111985 |
| sp. | MTR13495 | CTMZ6925 | Trancoso | BA | KM111810 | KM111901 | - | KM111986 |
| sp. | MTR13545 | CTMZ6927 | Trancoso | BA | KM111811 | KM111902 | - | KM111987 |
| sp. | MTR13547 | CTMZ6929 | Trancoso | BA | KM111812 | KM111903 | - | KM111988 |
| sp. | MTR13548 | CTMZ6930 | Trancoso | BA | KM111813 | KM111904 | - | KM111989 |
| sp. | MTR13565 | CTMZ6932 | Trancoso | BA | KM111814 | KM111905 | - | KM111990 |
| sp. | MTR13579 | CTMZ6933 | Trancoso | BA | KM111815 | KM111906 | - | KM111991 |
| sp. | MTR13589 | CTMZ6935 | Trancoso | BA | KM111816 | KM111907 | KM112020 | - |