Citation: Huemer P, Karsholt O, Mutanen M (2014) DNA barcoding as a screening tool for cryptic diversity: an example from Caryocolum, with description of a new species (Lepidoptera, Gelechiidae). ZooKeys 404: 91–111. doi: 10.3897/zookeys.404.7234
We explore the potential value of DNA barcode divergence for species delimitation in the genus Caryocolum Gregor & Povolný, 1954 (Lepidoptera, Gelechiidae), based on data from 44 European species (including 4 subspecies). Low intraspecific divergence of the DNA barcodes of the mtCOI (cytochrome c oxidase 1) gene and/or distinct barcode gaps to the nearest neighbor support species status for all examined nominal taxa. However, in 8 taxa we observed deep splits with a maximum intraspecific barcode divergence beyond a threshold of 3%, thus indicating possible cryptic diversity. The taxonomy of these taxa has to be re-assessed in the future. We investigated one such deep split in Caryocolum amaurella (Hering, 1924) and found it in congruence with yet unrecognized diagnostic morphological characters and specific host-plants. The integrative species delineation leads to the description of Caryocolum crypticum sp. n. from northern Italy, Switzerland and Greece. The new species and the hitherto intermixed closest relative C. amaurella are described in detail and adults and genitalia of both species are illustrated and a lectotype of C. amaurella is designated; a diagnostic comparison of the closely related C. iranicum Huemer, 1989, is added.
Lepidoptera, Gelechiidae, Caryocolum, species delineation, integrative taxonomy, DNA barcode, morphology, Europe
The genus Caryocolum Gregor & Povolný, 1954 is one of the most species-rich genera of European Gelechiidae (
Extensive generic descriptions and diagnoses of European species of Caryocolum have been published in several reviews, particularly
Specimens. Our study is based on about 50 specimens of the Caryocolum amaurella (Hering, 1924) species-group and an uncounted number of European Caryocolum, exceeding 1000 specimens, but only partially used for genetic analysis (see below). Most of the material was traditionally set and dried or alternatively spread; a few specimens are only pinned. Genitalia preparations followed standard techniques (Robinson 1976) adapted for male genitalia of Gelechiidae and (some) female genitalia of Caryocolum by the so-called “unrolling technique” (Pitkin 1986,
DNA Barcodes. Full-length lepidopteran DNA barcode sequences are a 648 base-pair long segment of the 5’ terminus of the mitochondrial COI gene (cytochrome c oxidase 1). DNA samples (dried leg) were prepared according to the accepted standards. Legs from 250 specimens of Caryocolum were processed at the Canadian Centre for DNA Barcoding (CCDB, Biodiversity Institute of Ontario, University of Guelph) to obtain DNA barcodes using the standard high-throughput protocol described in
Photographic documentation. Photographs of the adults were taken with an Olympus SZX 10 binocular microscope and an Olympus E 3 digital camera and processed using the software Helicon Focus 4.3 and Adobe Photoshop CS4 and Lightroom 2.3. Genitalia photographs were taken with an Olympus E1 Digital Camera from Olympus BH2 microscope.
BMNH The Natural History Museum (British Museum, Natural History) London (United Kingdom)
TLMF Tiroler Landesmuseum Ferdinandeum, Innsbruck, Austria
ZMUH Zoological Museum, University of Helsinki, Finland
ZMUC Zoological Museum, Natural History Museum of Denmark, Copenhagen, Denmark
ZMUO Zoological Museum, University of Oulu, Finland
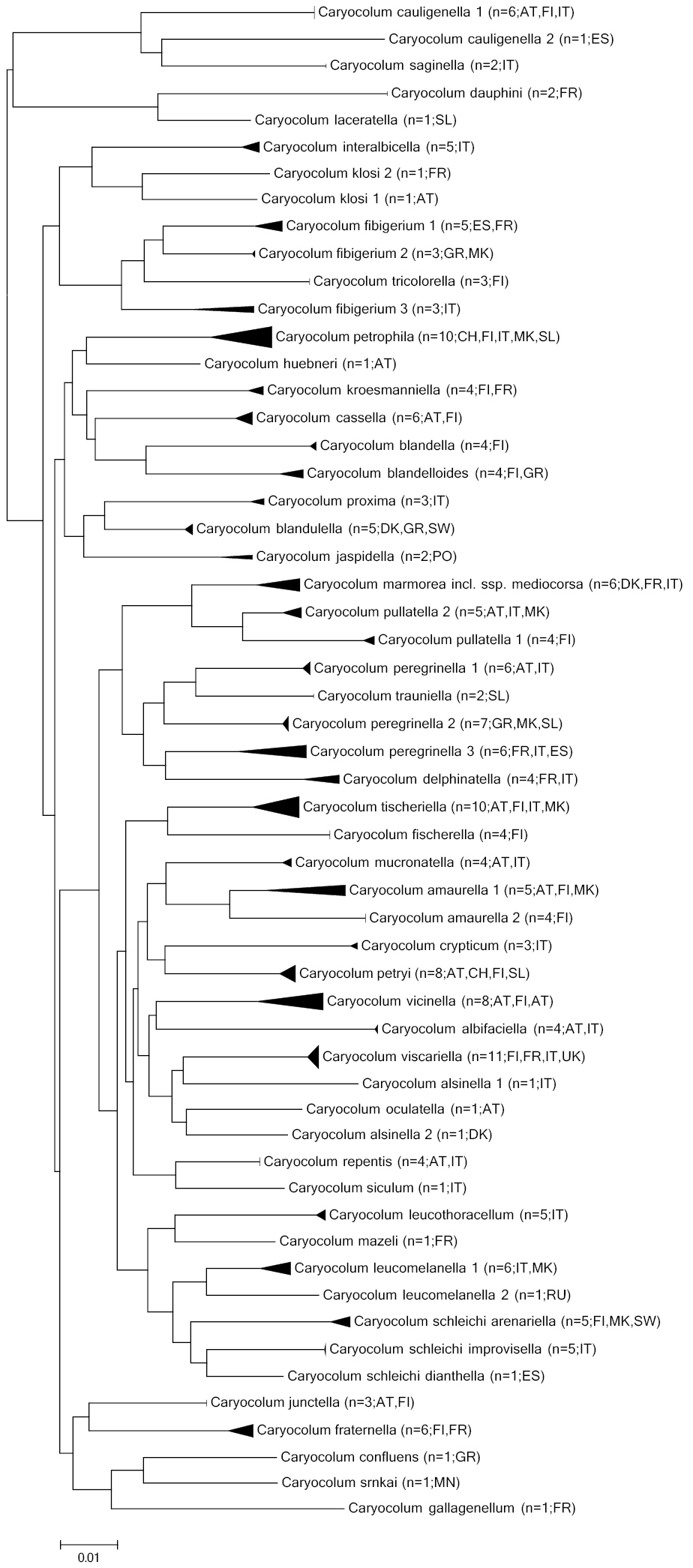
Forty-four of 51 European species were successfully sequenced, resulting in a full-length barcode fragment for 191 specimens and more than 500 bp for further 26 specimens (Fig. 1, Table 1, Suppl. material 1). Nine shorter sequences were not included in the analysis and sequencing of 24 specimens failed. The maximum intraspecific K2P distance varies from 0% in several species to 6.27% in Caryocolum fibigerium. Ten species have a high maximum intraspecific divergence greater than 2%. In six species (newly described species excluded) with a medium divergence greater than 3% potential cryptic diversity should be investigated. Furthermore, the intraspecific divergence of more than 3% in Caryocolum schleichi, a species separated into 3 allopatric subspecies, is beyond variation typically found within species, supporting their status as valid species. The only other subspecies we have examined are nominotypical Caryocolum marmorea and the recently separated Caryocolum marmorea mediocorsa with a very low divergence of 0.3%.
Neighbour-joining tree (Kimura 2 parameter, built with MEGA 5; cf.
Intraspecific mean K2P (Kimura 2 Parameter) divergences, maximum pairwise distances and distance to nearest neighbor.
| Species | Mean Intra-Sp | Max Intra-Sp | Nearest Neighbour | Nearest Species | Distance to NN |
|---|---|---|---|---|---|
| Caryocolum alsinella | 4.85 | 4.85 | PHLAE427-11 | Caryocolum oculatella | 3.81 |
| Caryocolum amaurella | 3.05 | 4.76 | LEATC402-13 | Caryocolum mucronatella | 5.21 |
| Caryocolum blandella | 0.16 | 0.3 | LEFIK150-10 | Caryocolum blandelloides | 5.78 |
| Caryocolum blandelloides | 0.4 | 0.81 | LEFIB755-10 | Caryocolum blandella | 5.78 |
| Caryocolum blandulella | 0.21 | 0.46 | LEATD656-13 | Caryocolum proxima | 3.94 |
| Caryocolum cassella | 0.42 | 0.61 | PHLAI019-12 | Caryocolum blandulella | 5.07 |
| Caryocolum cauligenella | 1.99 | 6.95 | PHLAA069-09 | Caryocolum saginella | 6.61 |
| Caryocolum confluens | N/A | N/A | PHLAF489-11 | Caryocolum srnkai | 4.54 |
| Caryocolum crypticum | 0.21 | 0.31 | LEATC-402-13 | Caryocolum mucronatella | 5.41 |
| Caryocolum dauphini | 0 | 0 | PHLAB900-10 | Caryocolum laceratella | 5.29 |
| Caryocolum delphinatella | 1.02 | 1.39 | PHLAI203-13 | Caryocolum marmorea mediocorsa | 4.57 |
| Caryocolum fibigerium | 3.4 | 6.27 | LEFIF467-10 | Caryocolum tricolorella | 4.67 |
| Caryocolum fischerella | 0 | 0 | LEFIC281-10 | Caryocolum tischeriella | 4.5 |
| Caryocolum fraternella | 0.47 | 1.7 | PHLAI156-12 | Caryocolum junctella | 4.55 |
| Caryocolum gallagenellum | N/A | N/A | PHLAI019-12 | Caryocolum blandulella | 6.54 |
| Caryocolum huebneri | N/A | N/A | LEFIJ1014-11 | Caryocolum petrophila | 4.88 |
| Caryocolum interalbicella | 0.4 | 0.77 | PHLAI156-12 | Caryocolum junctella | 5.55 |
| Caryocolum jaspidella | 1.08 | 1.08 | PHLAI019-12 | Caryocolum blandulella | 4.39 |
| Caryocolum junctella | 0 | 0 | LEFIF480-10 | Caryocolum fraternella | 4.55 |
| Caryocolum klosi | 4.25 | 4.25 | PHLAA055-09 | Caryocolum interalbicella | 5.56 |
| Caryocolum kroesmanniella | 0.31 | 0.61 | LEEUA184-11 | Caryocolum blandulella | 4.9 |
| Caryocolum laceratella | N/A | N/A | PHLAI447-13 | Caryocolum dauphini | 5.29 |
| Caryocolum leucomelanella | 1.47 | 3.79 | PHLAG331-12 | Caryocolum mazeli | 3.76 |
| Caryocolum leucothoracellum | 0.12 | 0.3 | PHLAG331-12 | Caryocolum mazeli | 4.24 |
| Caryocolum marmorea mediocorsa | 0 | 0 | LEEUA182-11 | Caryocolum marmorea | 0.3 |
| Caryocolum marmorea | 1 | 1.54 | PHLAI203-13 | Caryocolum marmorea mediocorsa | 0.3 |
| Caryocolum mazeli | N/A | N/A | LEATE421-13 | Caryocolum leucomelanella | 3.76 |
| Caryocolum mucronatella | 0.3 | 0.46 | PHLAE427-11 | Caryocolum oculatella | 4.87 |
| Caryocolum oculatella | N/A | N/A | LEEUA388-11 | Caryocolum alsinella | 3.81 |
| Caryocolum peregrinella | 3.58 | 5.69 | PHLAB899-10 | Caryocolum trauniella | 3.93 |
| Caryocolum petrophila | 0.97 | 2.26 | PHLAH147-12 | Caryocolum huebneri | 4.88 |
| Caryocolum petryi | 0.23 | 0.61 | PHLAD576-11 | Caryocolum repentis | 3.85 |
| Caryocolum proxima | 0.41 | 0.61 | PHLAI019-12 | Caryocolum blandulella | 3.94 |
| Caryocolum pullatella | 2.07 | 3.61 | LEATC292-13 | Caryocolum marmorea | 3.12 |
| Caryocolum repentis | 0 | 0 | PHLAE429-11 | Caryocolum siculum | 3.33 |
| Caryocolum saginella | 0 | 0 | LEFIJ778-10 | Caryocolum cauligenella | 6.61 |
| Caryocolum schleichi dianthella | N/A | N/A | PHLAD573-11 | Caryocolum schleichi improvisella | 3.42 |
| Caryocolum schleichi improvisella | 0.06 | 0.15 | PHLSA085-11 | Caryocolum schleichi dianthella | 3.42 |
| Caryocolum schleichi arenariella | 0.77 | 1.24 | PHLSA085-11 | Caryocolum schleichi dianthella | 3.74 |
| Caryocolum siculum | N/A | N/A | PHLAD576-11 | Caryocolum repentis | 3.33 |
| Caryocolum srnkai | N/A | N/A | PHLAG580-12 | Caryocolum confluens | 4.54 |
| Caryocolum tischeriella | 1.09 | 2.02 | PHLAD576-11 | Caryocolum repentis | 4.01 |
| Caryocolum trauniella | 0 | 0 | PHLAB622-10 | Caryocolum peregrinella | 3.93 |
| Caryocolum tricolorella | 0 | 0 | PHLAI014-12 | Caryocolum fibigerium | 4.67 |
| Caryocolum vicinella | 1.48 | 2.7 | PHLAF105-11 | Caryocolum leucomelanella | 5.36 |
| Caryocolum viscariella | 0.22 | 0.47 | LEEUA388-11 | Caryocolum alsinella | 4.16 |
Sequences of the COI barcode region of all analysed morphospecies reveal significant interspecific genetic distances with barcode gaps ranging from a minimum of 3.11% to the nearest neighbour (Caryocolum pullatella – Caryocolum marmorea) to a maximum of 6.61% (Caryocolum saginella – Caryocolum cauligenella).
The Caryocolum amaurella species-group as defined by
Caryocolum Gregor & Povolný, 1954: 87.
Type species. Gelechia leucomelanella Zeller, 1839: 138.
http://zoobank.org/5E1FB9E5-3A65-49C6-80BF-A5CA7C4FFF99
http://species-id.net/wiki/Caryocolum_crypticum
Figs 2–3, 6–7, 10–11, 14–15Holotype: ♀ (Fig. 2), Italia sept., Teriolis merid., Laatsch, 1000 m, 29.6.1987 e.l. (Silene otites 10.5.), leg. Huemer, slide GEL 1234 ♀ (TLMF).
Paratypes. Italy: 1 ♂, South Tyrol, Vinschgau, Schleiser Leiten, 1350 m, 6.7.2013, leg. Huemer, slide GEL 1215, dna barcode id TLMF Lep 12313 (TLMF); 1 ♂ [without abdomen], same data (TLMF); 1 ♀, same data, but 18.8.2013, slide GEL 1232, dna barcode id TLMF Lep 11883 (TLMF); 1 ♀, same data, but dna barcode id TLMF Lep 11882 (TLMF); 1 male [without abdomen], 8 ♀, same data, but 7.9.2013 (TLMF); 1 ♂, South Tyrol, Taufers, 1300 m, 22.8.1978, leg. Burmann, slide GU 86/041 P. Huemer (TLMF). Switzerland: 2 ♀, Wallis, Martigny-Rosel, 460 m, 28.6.–14.7.1983 e.l. (Silene otites), leg. Whitebread (Naturhistorisches Museum Basel, Switzerland). Greece: 1 ♀, Larisa, Ossa Oros, 1.5 km N Spilia, 940 m, 13.6.1988 e.l. (Silene nutans), leg. Huemer (TLMF).
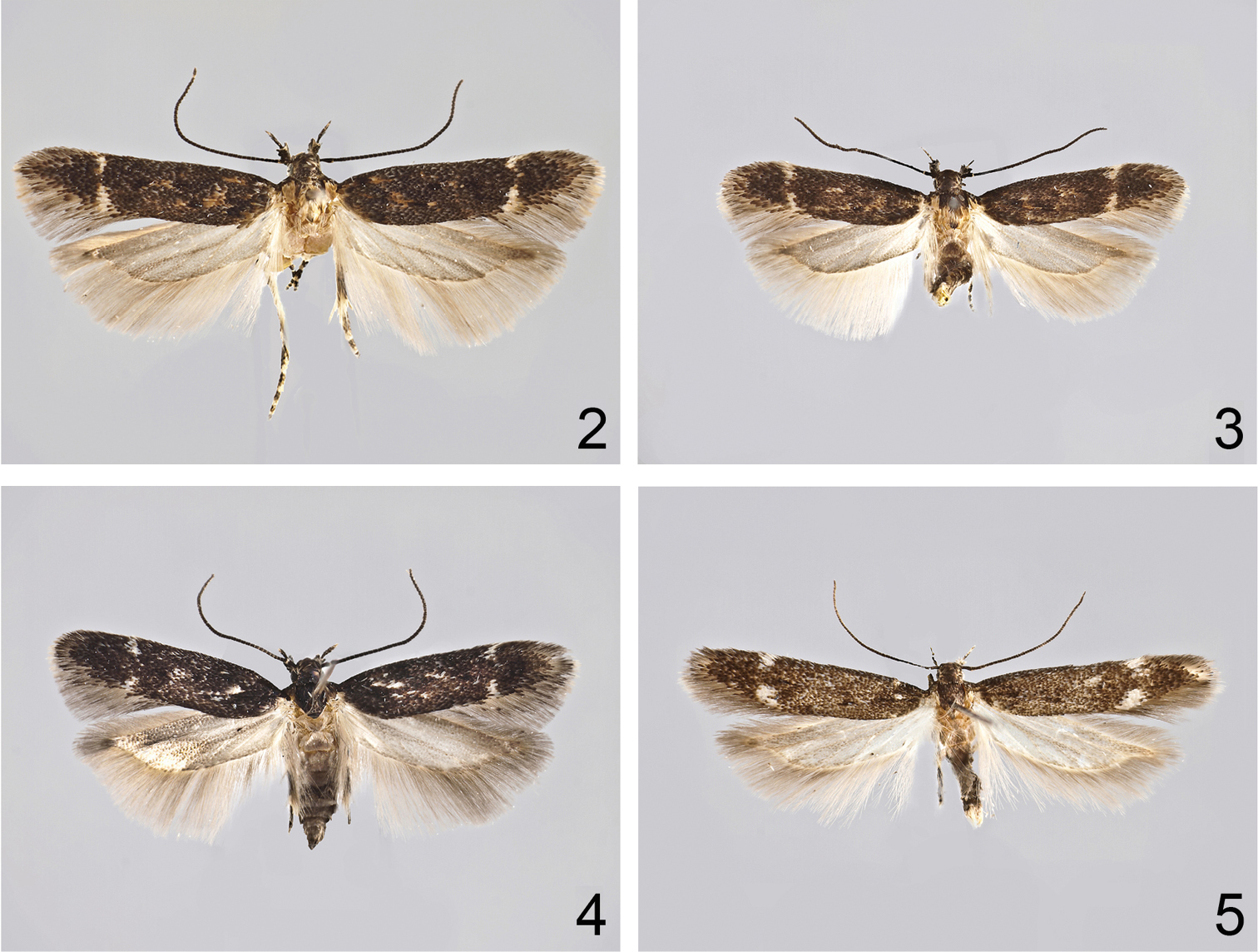
Adults. 2 Caryocolum crypticum sp. n., holotype 3 Caryocolum crypticum sp. n., paratype, female, Greece 4 Caryocolum amaurella, male, Finland 5 Caryocolum amaurella, male, Austria.
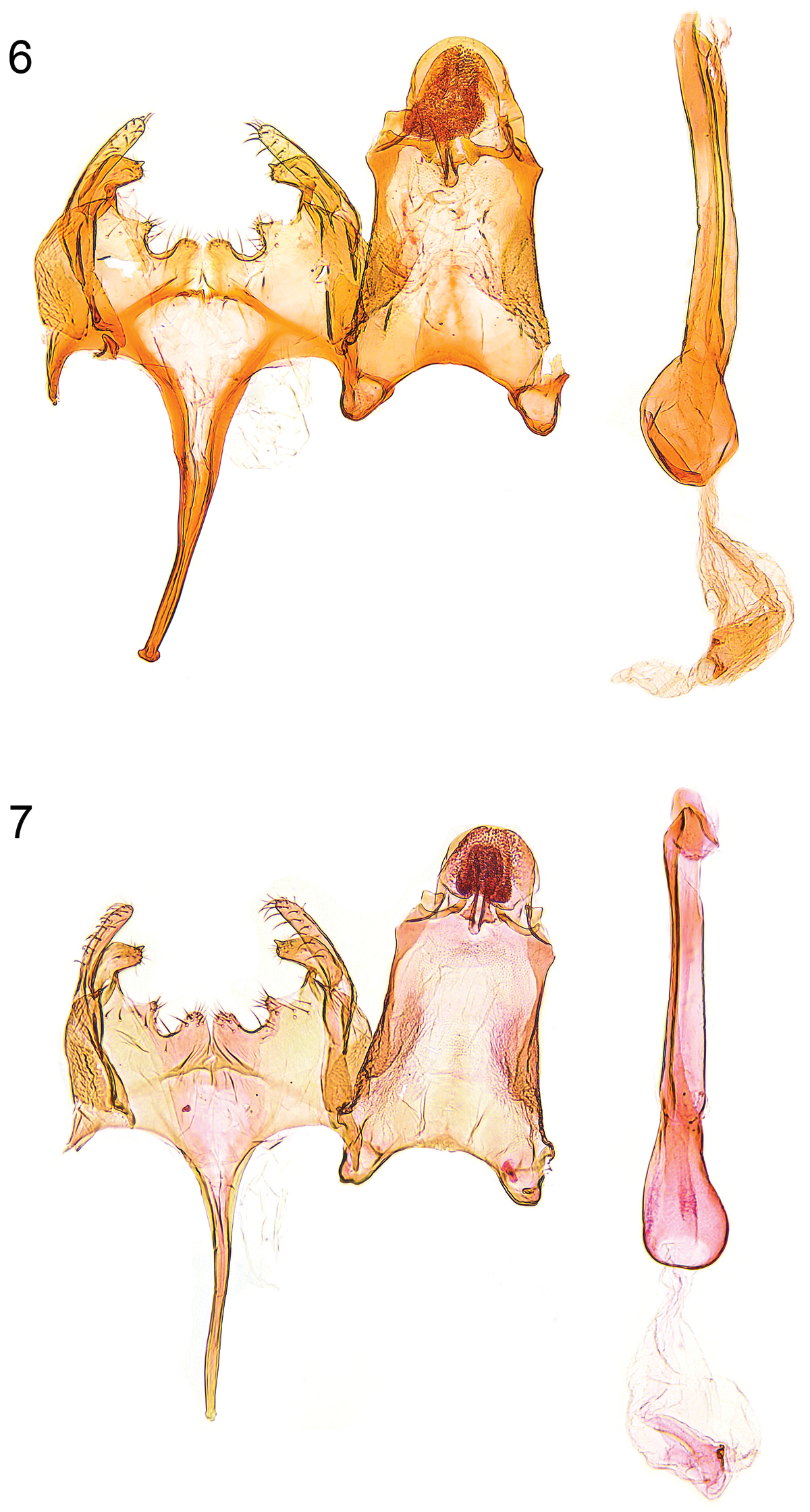
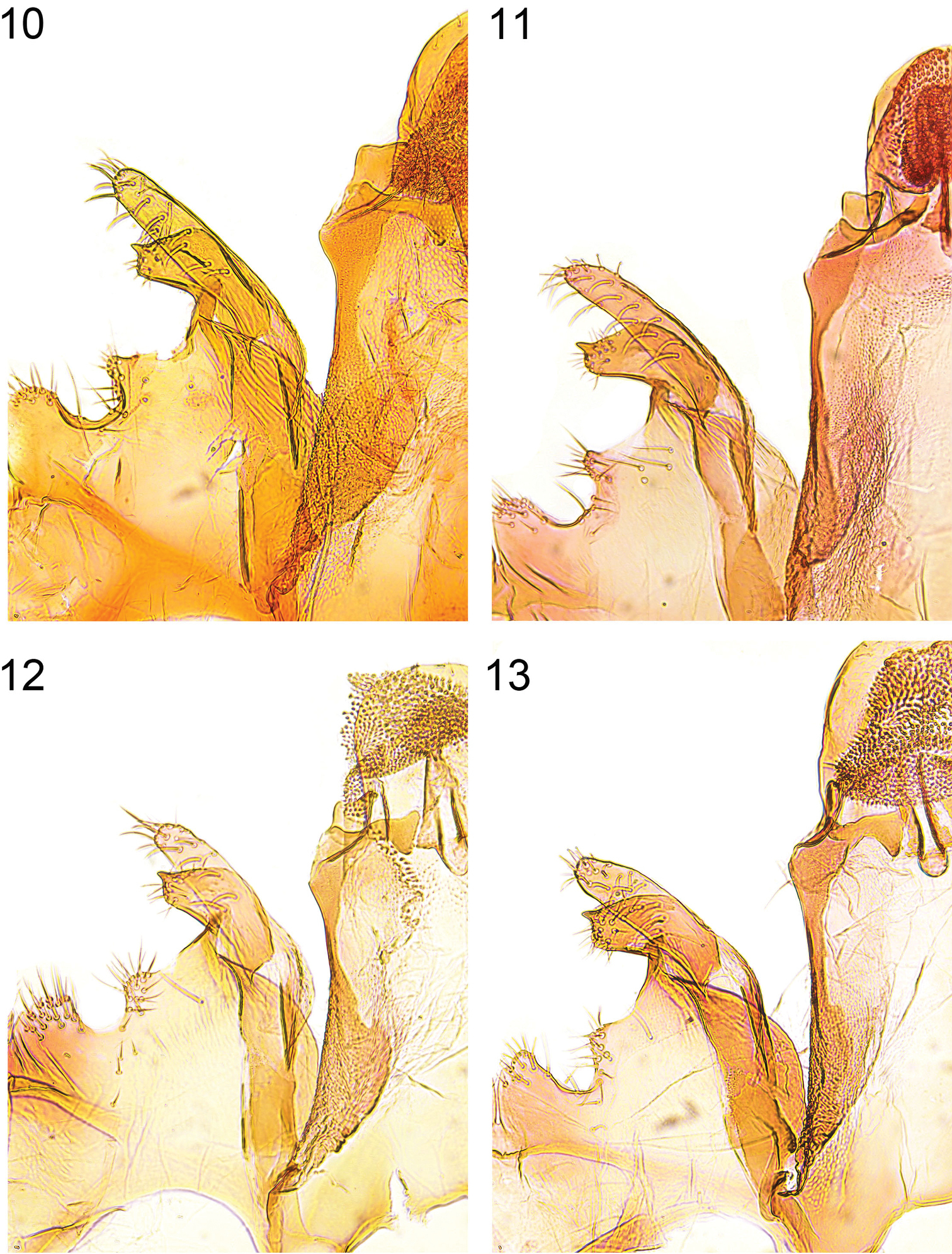
Caryocolum crypticum sp. n. is externally similar to several other species of the genus and can be best recognized by the largely unmarked forewings with cream costal and tornal spots. From its closest relatives Caryocolum amaurella and Caryocolum iranicum it differs by the rusty brown distal half of the thorax and the concolorous tegulae, the dark brown forewings with rusty brown scales, and the cream colours of the costal and tornal spots. The male genitalia of Caryocolum crypticum are very similar to those of Caryocolum amaurella but the valva is more slender and slightly longer (see Figs 6–7, 10–11 versus 8–9, 12–13). The similar Caryocolum iranicum differs by the shape of the sacculus with almost straight dorsal margin (see
Male genitalia. 6 Caryocolum crypticum sp. n., paratype, Italy, slide GU 86/041 P.Huemer 7 Caryocolum crypticum sp. n., paratype, Italy, slide GEL 1215 P.Huemer.
Adult (Figs 2–3). Wingspan 10.5-14 mm. Segment 2 of labial palpus with a few cream-coloured scales on inner and upper surface, blackish brown on outer and lower surface; segment 3 almost black with light tip. Antenna black, indistinctly lighter ringed. Head with light yellow frons and black neck; thorax blackish brown with rusty brown posterior part; tegulae rusty brown except for blackish brown base. Forewing blackish brown, mottled with some rusty brown, particularly in proximal half; supplementary black spots in fold and in cell obscure; costal and tornal spot small, cream, separated. Hindwing light grey.
Variation. No variation observed except for size, which differs considerably in two reared specimens from Italy and Greece.
Male genitalia (Figs 6–7, 10–11). Uncus subovate; tegumen stout; transtilla membranous; valva moderately short and slender, digitate, apex rounded; sacculus short, with angular ventral and weakly convex dorsal margin, apically pointed; posterior margin of vinculum with deep medial emargination and slight medial incision, two pairs of short processes developed; saccus long, comparatively broad at base, distal part gradually tapered; phallus long and slender, weakly curved, with some minute cornuti apically.
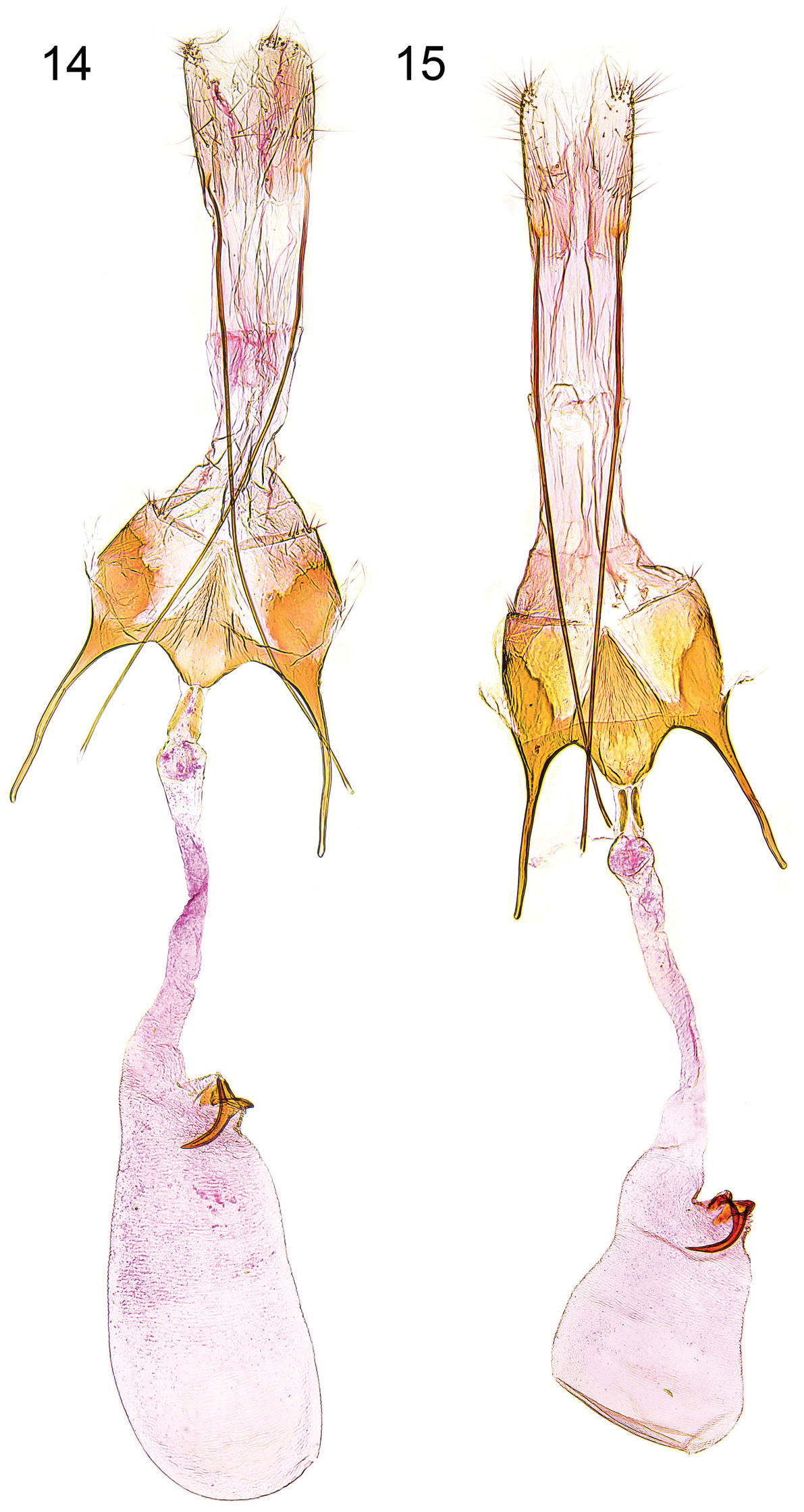
Female genitalia (Figs 14–15). Segment VIII without processes, subgenital plate sub-triangular, with numerous narrow folds, separated from sclerotized lateral plates by membranous zone; apophysis anterior about length of segment VIII; antrum short, about one quarter length of apophysis anterior, nearly cup-shaped; posterior part of ductus bursae with pair of short sclerites, extending to middle of apophysis anterior, and with two tiny sclerites anteriorly; signum with crescent-shaped base, long and slender, strongly bent hook.
Molecular data. The intraspecific divergence of the barcode region is low with mean intraspecific divergence of 0.21% and maximum intraspecific divergence of 0.31% (n=3). The distance to the nearest neighbour Caryocolum mucronatella is 5.41%, the divergence to the morphologically closest Caryocolum amaurella is 6.82%.
The name “crypticum” refers to the cryptic morphology of the species and is derived from the latinized adjective crypticus.
The species is known from widely separated localities in northern Italy, Switzerland and Greece, indicating a more widespread distribution in Sub-Mediterranean and Mediterranean Europe. However, the host-plants are much more widespread, ranging to northern Europe in the north and to Central Asia in the east. No sympatric occurrence with Caryocolum amaurella is reported though the two taxa can occur close to one another in the Alps.
The larva has been found in early spring, feeding in the stem of Silene otites (L.) Wibel (Caryophyllaceae) (
The majority of collected material belongs to females whereas Caryocolum amaurella is mainly known from the male sex. This may indicate differences in attraction to artificial lights or a female-biased sex ratio in Caryocolum crypticum.
http://species-id.net/wiki/Caryocolum_amaurella
Figs 4–5, 8–9, 12–13, 16–17Lectotype ♂[with nine labels]: ‘Fennia Ab Bromarf’ ‘R. F:tius’ ‘21.7.21’ [piece of celluloid where genitalia was mounted] ‘Type ♂’ [red] ‘Lita amaurella m. det. Mart. Hering ♂’ ‘Mus. Zool. H:fors spec. typ. No 7016 Lita amaurella Hering’ ‘Lita amaurella m. ♂ Sch.-Armatur Bromarf 21.7.21 Fabritius’ ‘LECTOTYPE O. Karsholt design.’.
Finland: 1 ♂, Ab, Naantali, 25.8.1965, leg. Karvonen, slide Karsholt 2719; 2 ♂, N, Ekenäs, 17.–20.7.1980, leg. Fibiger; 1 ♂, N, Helsinki, 25.7.1982, leg. Schnack; 1 ♂, N, Borgå lk., Tirmo, 19–20.7.1980, leg. Fibiger; 5 ♂, same data, but 1.–2.8.1982, leg. Schnack; 2 ♂, U, Sluntle, 18.–31.7.1982, leg. Karsholt (all ZMUC); 6 ♂, 5 ♀, U, Porvoo, 6698:3426 Ånäs, e.l. 2012 (Lychnis viscaria), leg. Hirvonen (ZMUO); 4 ♂, V, Dragsfjärd, 664:3249, 2008, leg. Mutanen & Välimäki (ZMUO); 1 ♂, U, Hanko, 6642:3289, 2007, leg. Mutanen & Välimäki (ZMUO). Sweden: 2 ♂, Sk, Maglehen, 10.7.1965, leg. Svensson (TLMF, ZMUC); 1 ♂, Sm, Högsby, 13.7.1968, leg. Johansson; 1 ♂, Öl, Ödeshög, 17.7.1972, leg. Karsholt, slide Karsholt 1806; 1 ♂, St. Alvar, Tornrör, 25.7.1997, leg. Hendriksen, slide Hendriksen 1953; 2 ♂, Öl, Gårdby, 2.8.1999, leg. Hendriksen, slide Hendriksen 2411, 2415; 1 ♂, same data, but 22.7.2000; 1 ♂, Gtl., Hejnum Häller, 30.7.1977, leg. Hendriksen, slide Hendriksen 1944; 1 ♂, Ög, Ödeshög, 17.7.1972, leg. Karsholt; Upl., Film, 12.7.1995, leg. Hendriksen (all ZMUC). Norway: 2 ♂, On, Vinstra, 19.–29.7.1983, leg. Karsholt & Michelsen, slide Karsholt 4294, 4295; 2 ♀, same data, but 4.–5.7.1987, leg. Karsholt, slide Hendriksen 2099; 2 ♂, same data, but 9.8.1996, leg. Hendriksen (all ZMUC). Denmark: 1 ♂, Bornholm, Rø, 7.1892, leg. Gudmann, slide Wolff 2593; 5 ♂, 1 ♀, same data, but 28.7.1978, leg. Schnack, slide Schnack 1118; 1 ♂, Bornholm, Gudhjem, 1 ♂, 2 ♀, 29.6–3.7.1920, leg. Gudmann, slide Wolff 2625, 3682; 1 ♂, 1 ♀, same data, but e.l. 5.1921 (Lychnis viscaria), bred 21. & 28.6.1921, leg. Gudmann, slide Wolff 3681(all ZMUC); 6 ♂, 8 ♀, Bornholm, Hammeren, 18.7.1977, leg. Karsholt & Schnack, slide Hendriksen 1767, Karsholt 2948 TLMF, ZMUC); 2 ♂, same data, but 25.7.1977, leg. Schnack; 4 ♂, same data, but 16. –25.7.1978, leg. Schnack; 4 ♂, same data, but 19.–22.7.1979, leg. Hendriksen; 6 ♂, same data, but 29 –30.7.1981, leg. Hendriksen, slide Hendriksen 385, 561, 722; Bornholm, Randkløve, 1 ♂, 22.7.1977, leg. Schnack; Bornholm, 1 ♀, Ringe Bakker, 16.7.1978, leg. Schnack (all ZMUC). Germany: 1 ♂, 1 ♀, Lausitz, Umg. Bautzen, e.l. 1935 (Lychnis viscaria), leg. Starke (BMNH); 2 ♂, Thüringen, Bad Blankenburg, 14.7.1964, leg. Steuer (TLMF); 1 ♂, Thüringen, Bad Blankenburg, 8.7.1972, leg. Steuer (TLMF). France: 2 ♂, Alpes Maritimes, Col de la Cayolle, 2200–2300 m, 29.–30.7.2005, leg. Skou, slide Hendriksen 5364 (ZMUC). Austria: 1 ♂, Niederösterreich, Jauerling, 24.7.1935 (TLMF); 2 ♂, Oberösterreich, Windischgarsten, Veichltal, 23.7.1976, leg. Wimmer (TLMF); 1 ♂, Oberösterreich, Waldhausen, Schwarzenberg, 6.8.1997, leg. Wimmer (TLMF); 9 ♂, Kärnten, St. Jakob im Lesachtal, Mussen E, 1680–1800 m, 4.8.1999, leg. Huemer & Erlebach (TLMF). Slovakia: 1 ♀, Pol’ana, 28.7.1989, leg. Patocka (ZMUC). Macedonia: 4 ♂, NP Mavrovo, Korab, Korabska jezero, Kobilino pole, 2080–2180 m, 28.7.–1.8.2011, leg. Huemer & Tarmann (TLMF). Turkey: 2 ♂, 1 ♀, prov. Sivas, 10 km W Görün, 1650 m, 27.7.1989, leg. Esser & Fibiger, slide Huemer GU 90/130, GU 91/215; 4 ♂, prov. Erzerum, Kop Pass, 1750 m, 15.–16.9.1993, leg. Fibiger, slide Hendriksen 2889, 2894; 1 ♂, prov. Erzincan, Kizildaĝ, Geçidi, 2100 m, 19.8.1993, leg. Schepler, slide Hendriksen 2384 (all ZMUC).
See above.
Adult (Figs 4–5). Wingspan 10–14 mm. Segment 2 of labial palpus bone-white on inner and upper surface, blackish grey on outer and lower surface; segment 3 almost black with light tip. Antenna black, indistinctly lighter ringed. Head with light yellow frons and black neck; thorax and tegula black mottled with brown. Forewing blackish grey mottled with some light brown; base black; two indistinct black spots in fold; one oblique spot above it and one in cell; some white scales before and after these spots; costal and tornal spot small, white, rarely fused. Hindwing light grey.
Variation. The colour of the forewings varies from greyish to blackish. Worn specimens look lighter than fresh ones. Sometimes there are no white scales in the middle of the wing.
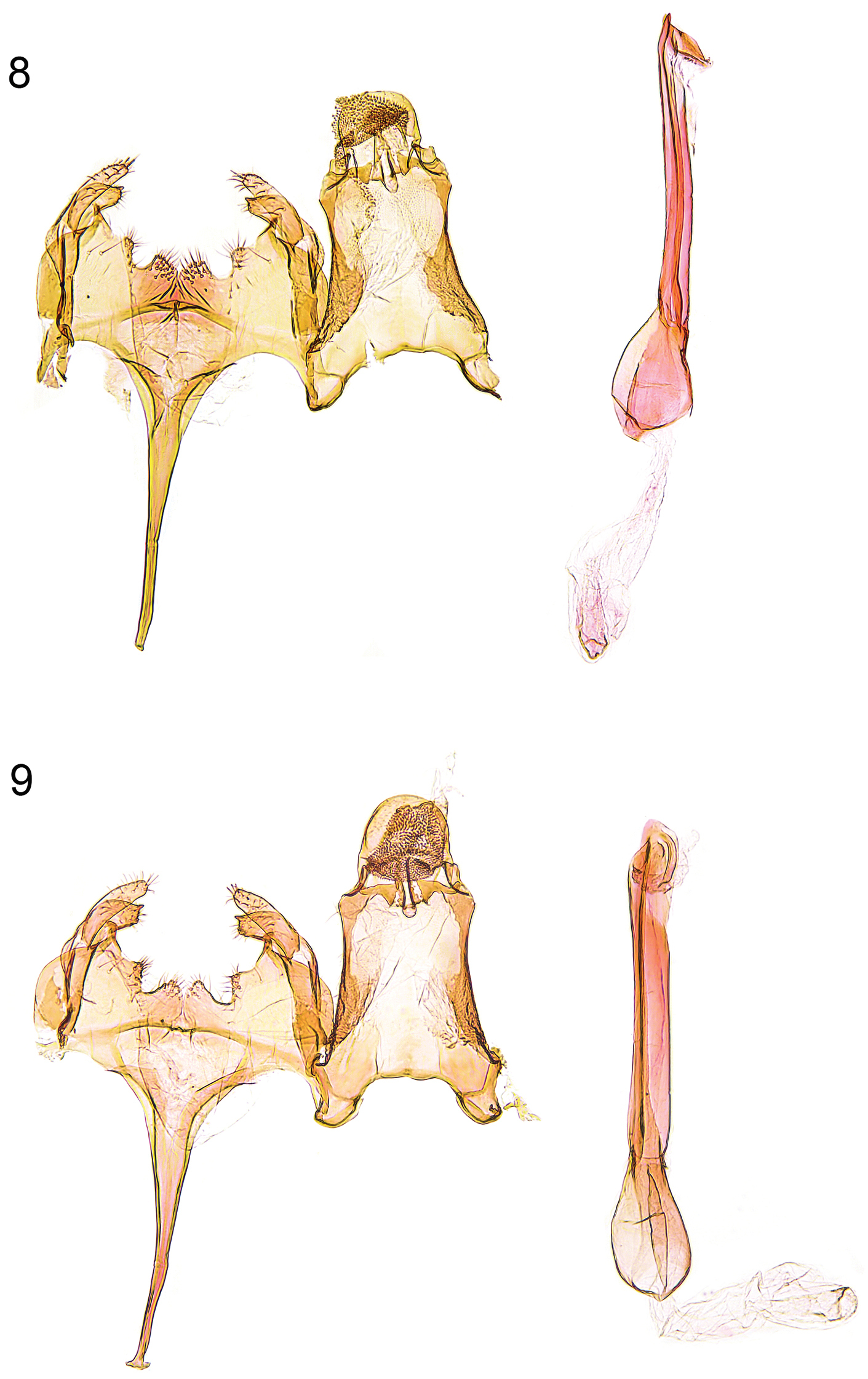
Male genitalia (Figs 8–9, 12–13). Uncus subovate; tegumen stout; transtilla membranous; valva short, moderately stout, apex rounded; sacculus short, with angular ventral and convex dorsal margin, apically pointed; posterior margin of vinculum with deep medial emargination and slight medial incision, two pairs of short processes developed; saccus long, comparatively broad at base, distal part gradually tapered; phallus long and slender, weakly curved, with some minute cornuti apically.
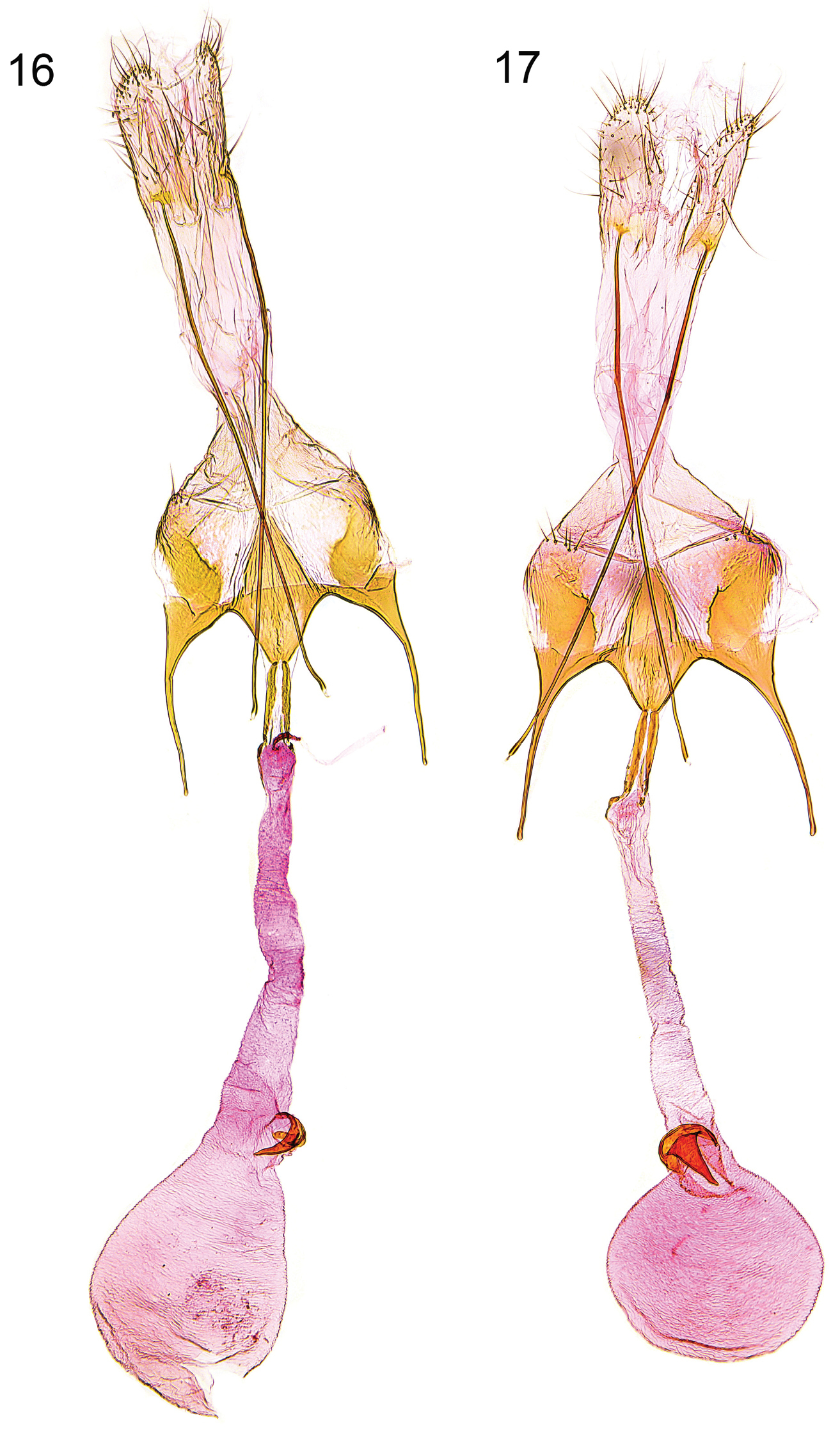
Female genitalia (Figs 16–17). Segment VIII without processes, subgenital plate sub-triangular, with numerous narrow folds, separated from sclerotized lateral plates by membranous zone; apophysis anterior slightly longer than segment VIII; antrum moderately short, about one-third to one-quarter length of apophysis anterior, broadly funnel-shaped; posterior part of ductus bursae with pair of lateral sclerites, extending to anterior third of apophysis anterior, and with two tiny sclerites anteriorly; signum with crescent-shaped base, short and stout, strongly bent hook.
Male genitalia. 8 Caryocolum amaurella (Hering), Finland, slide GU 14/1373 P.Huemer; 9 Caryocolum amaurella, Finland, slide GU 14/1374 P.Huemer.
Details of male genitalia (vinculum-valva-complex). 10 Caryocolum crypticum sp. n., paratype, Italy, slide GU 86/041 P.Huemer 11 Caryocolum crypticum sp. n., paratype, Italy, slide GEL 1215 P.Huemer 12 Caryocolum amaurella, Finland, slide GU 14/1373 P.Huemer 13 Caryocolum amaurella, Finland, slide GU 14/1374 P.Huemer.
Female genitalia. 14 Caryocolum crypticum sp. n., holotype, slide GEL 1234 P.Huemer 15 Caryocolum crypticum sp. n., paratype, Italy, slide GEL 1232 P.Huemer.
Female genitalia. 16 Caryocolum amaurella, Finland, slide GU 14/1372 P.Huemer 17 Caryocolum amaurella, Finland, slide GU 14/1371 P.Huemer.
Molecular data. The intraspecific divergence of the barcode region is high with mean intraspecific divergence of 3.01% and maximum intraspecific divergence of 4.62% (n=9). The distance to the nearest neighbour Caryocolum mucronatella is 5.21%, the divergence to the morphologically closest Caryocolum crypticum is 6.82%. The extraordinary high intraspecific divergence with 4 haplotypes is partially related to geographical pattern. However, we also found two haplotypes within one population in Finland and morphology does not support cryptic diversity.
With certainty known from scattered records from northern and Central Europe and Turkey. All the specimens from north of the Alps that we have been able to cross-check are correctly attributed to Caryocolum amaurella. However, recent records from Ukraine (
Bionomics. The larva has been recorded feeding on Silene viscaria (L.) Jess (= Lychnis viscaria L. (Caryophyllaceae) (
Lita amaurella was described from an unspecified number of specimens of both sexes (‘♂, ♀’) from Finland (Bromarf) (
Turkish specimens of Caryocolum amaurella examined by us differ from European specimens of this species by the thorax with rusty brown posterior part and the rusty brown tegulae with blackish brown base, similar to Caryocolum crypticum, and they are thus hardly separable from the latter on external characters. The genitalia of both sexes of Caryocolum amaurella from Turkey agree in all details with those of European Caryocolum amaurella and, because no contradicting genetic data is currently available, we consider them as belonging to that species.
One of the examined specimens of Caryocolum amaurella from Turkey was collected in the same locality (Kizildaĝ Geçidi, prov. Erzincan) as a specimen Caryocolum iranicum in ZMUC. The latter species, which is only known from a few specimens, differs, as stated above, in characters of the male genitalia.
The genus Caryocolum is a rare example of European Microlepidoptera which has gained significant attention from specialists during the last decades. Several monographic papers, from
Intraspecific variation shows a different pattern. The majority of species has a low (<2%) maximum intraspecific divergence and thus seems taxonomically well defined. However, a remarkable number of species (8 species, nearly one quarter of all, 9 species with only one sample not considered) is characterized by maximum divergence exceeding 3% (Fig. 1). Such deep intraspecific splits often suggest the possibility of cryptic diversity (for examples in Lepidoptera, see
We are particularly grateful to Paul Hebert and his team at the Canadian Centre for DNA Barcoding (Guelph, Canada), whose sequencing work was enabled by funding from the Government of Canada to Genome Canada through the Ontario Genomics Institute. We are also grateful to the Ontario Ministry of Research and Innovation and to NSERC for their support of the BOLD informatics platform.
Stefan Heim (TLMF) is acknowledged for his kind assistance with photographic work. We thank Robert J. Heckford (Plymouth, GB), Petri Hirvonen (Porvoo, Finland), Thierry Varenne (Nice, France), Christian Wieser (Klagenfurt, Austria) and Josef Wimmer (Steyr, Austria) for providing material for our examination, and Lauri Kaila (ZMUH) for access to the type of Lita amaurella.
We are particularly indebted to the Promotion of Educational Policies, University and Research Department of the Autonomous Province of Bolzano - South Tyrol for helping to fund the project “Genetic biodiversity archive - DNA barcoding of Lepidoptera of the central Alpine region (South, East and North Tyrol)”. Furthermore fundings from inatura Erlebnis Naturschau (Dornbirn, Austria) are acknowledged.
Last, but not least, we thank Martin Corley (Faringdon, UK) for linguistic improvement of the manuscript.
Sample information for specimens included in this study.
Authors: Peter Huemer, Ole Karsholt, Marko Mutanen
Data type: species data
Explanation note: Process IDs are sequence identifiers in BOLD; Sample IDs are specimen identifiers; BINs are Barcode Identification Numbers in BOLD. Details of collecting data, images, sequences, and trace files for the barcoded specimens are available in the public BOLD dataset “DS-LECARY”, accessed at dx.doi.org/10.5883/DS-LECARY.
Copyright notice: This dataset is made available under the Open Database License (http://opendatacommons.org/licenses/odbl/1.0/). The Open Database License (ODbL) is a license agreement intended to allow users to freely share, modify, and use this Dataset while maintaining this same freedom for others, provided that the original source and author(s) are credited.