(C) 2013 Erika Espinoza. This is an open access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC-BY), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
Citation: Espinoza E, DuPont A, Valdés Á (2013) A tropical Atlantic species of Melibe Rang, 1829 (Mollusca, Nudibranchia, Tethyiidae). ZooKeys 316: 55–66. doi: 10.3897/zookeys.316.5452
A new species of Melibe is described based on two specimens collected in Florida. This new species is well differentiated morphologically and genetically from other species of Melibe studied to date. The four residue deletions in the cytochrome c oxidase subunit 1 protein found in all previously sequenced tropical species of Melibe sequenced (and Melibe rosea) are also present in this new species. These deletions do not appear to affect important structural components of this protein but might have fitness implications. This paper provides the first confirmed record of Melibe in the tropical western Atlantic Ocean.
new species, molecular taxonomy, anatomy, Opistobranchia, western Atlantic
Melibe Rang, 1829 (family Tethyiidae Rafinesque, 1815) is an unusual genus of cladobranch nudibranchs that feed by expanding a large oral hood fringed with sensory tentacles to capture small crustaceans. The digestive morphology of this group is largely modified, all species lack a radula and have a circularized stomach (
Melibe is also unusual biogeographically as it appears to be completely absent from the tropical Eastern Pacific and is poorly represented in the Atlantic Ocean. The only two confirmed records from the Atlantic are the two South African species Melibe rosea and Melibe liltvedi, which are found on the Atlantic side of the Cape Peninsula. Melibe viridis has been reported from the Mediterranean – originally as Melibe fimbriata (Alder & Hancock, 1864), but is considered a non-native species (
In this paper we describe the first species of Melibe from the tropical Atlantic Ocean based on two specimens recently collected in Florida. Molecular and morphological data obtained from the two specimens are compared with other congeners.
Two specimens were collected by SCUBA diving in Lake Worth Lagoon Florida, photographed alive, and preserved in pharmacy grade ethyl alcohol. Once in the laboratory they were transferred to ethanol 95%. All the specimens examined are deposited at the Natural History Museum of Los Angeles County, USA (abbreviated LACM).
The anterior portion of the digestive system and the reproductive system were dissected and drawn under a dissecting microscope with a camera lucida attachment. The stomach was also dissected to expose the stomach plates. The buccal mass was dissolved in a NaOH 10% solution to isolate the jaws. The jaws and the stomach were rinsed in distilled water, dried, mounted on a stub, and sputter coated for examination under a scanning electron microscope (SEM) Hitachi S-3000N.
DNA extraction was performed using a hot Chelex® protocol. Approximately 1-3 mg of the foot was cut into fine pieces for extraction. For the Chelex® extraction, the foot tissue was rinsed and rehydrated using 1.0 mL TE buffer (10 mM Tris, 1 mM EDTA, pH 8.0) for 20 minutes. A 10% (w/v) Chelex® 100 (100-200 mesh, sodium form, Bio-Rad) solution was prepared using TE buffer. After rehydration, the tissue mixture was then centrifuged, 975.00 μL of the supernatant was removed, and 175.00 μL of the Chelex® solution was added. Samples were then heated in a 56°C water bath for 20 minutes, heated in a 100°C heating block for 8 minutes, and the supernatant was used for PCR.
Histone-3 universal primers (H3 AF 5’-ATGGCTCGTACCAAGCAGACGGC-3’, H3 AR 5’-ATATCCTTGGGCATGATGGTGAC-3’ developed by
The master mix was prepared using 34.75 μL H2O, 5.00 μL Buffer B (ExACTGene, Fisher Scientific), 5.00 μL 25 mM MgCl2, 1.00 μL 40mM dNTPs, 1.00 μL 10mM primer 1, 1.00 μL primer 2, 0.25 μL 5 mg/mL Taq, and 2.00 μL of extracted DNA. Reaction conditions for H3 (universal) and 16S (universal) were as follows: lid heated to 105°C and initial denaturation of 94°C for 2 min, 35 cycles of 94°C for 30 s, 50°C for 30 s, and 72°C for 1 min, followed by a final elongation of 72°C for 7 min. Reaction conditions for CO1 (universal) were as follows: lid heated to 105°C and initial denaturation of 95°C for 3 min, 35 cycles of 94°C for 45 s, 45°C for 45 s, and 72°C for 2 min, followed by a final elongation step of 72°C for 10 min. PCR products yielding bands of appropriate size were purified using the Montage PCR Cleanup Kit (Millipore). Cleaned PCR samples were quantified using a NanoDrop 1000 Spectrophotometer (Thermo Scientific). Each primer was diluted to 4.0 pmol/μL to send out for sequencing with the PCR products. PCR products were diluted to 6.0, 7.5, and 11.5ng/μL for H3, 16S, and CO1 respectively. Samples were sequenced at Eton Bioscience, Inc. (San Diego, CA).
Sequences were deposited in GenBank (http://www.ncbi.nlm.nih.gov/genbank/) with the accession numbers KC992314 for CO1, KC992313 for 16S, and KC992315 for H3. Sequences of other species of Melibe and Tethys fimbria Linnaeus, 1767 were downloaded from GenBank (Table 1) and included in the phylogenetic analysis. Sequences for each gene were assembled and edited using GENEIOUS Pro 4.7.4 (
Specimens sequenced, including locality information, collection voucher numbers and GenBank accession numbers.
| Species | Voucher | Locality | GenBank accession numbers | ||
|---|---|---|---|---|---|
| CO1 | 16S | H3 | |||
| Tethys fimbria | - | - | AY345035 | AY345035 | EF133468 |
| Melibe leonina | LACM174849 | California, USA | GQ292059 | GU339202 | - |
| Melibe digitata | CASIZ175724 | Philippines | JX306069 | JX306061 | JX306076 |
| Melibe digitata | CASIZ177478 | Philippines | HM162699 | HM162617 | HM162523 |
| Melibe viridis | CASIZ176981 | Mozambique | JX306075 | JX306068 | JX306083 |
| Melibe viridis | CASIZ177524 | Philippines | HM162700 | HM162618 | HM162524 |
| Melibe rosea | CASIZ175734 | South Africa | JX306070 | JX306063 | JX306078 |
| Melibe rosea | CASIZ176355 | South Africa | JX306071 | JX306064 | JX306079 |
| Melibe rosea | CASIZ176367 | South Africa | JX306073 | JX306066 | JX306081 |
| Melibe rosea | CASIZ176356 | South Africa | JX306072 | JX306065 | JX306080 |
| Melibe rosea | CASIZ176392 | South Africa | HM162701 | HM162620 | HM162526 |
| Melibe engeli | CASIZ177625 | Philippines | - | HM162619 | HM162525 |
| Melibe engeli | CASIZ177757 | Philippines | - | JX306062 | JX306077 |
| Melibe arianeae | LACM3259 | Florida, USA | KC992314 | KC992313 | KC992315 |
To assess whether H3, 16S, and CO1 have significantly conflicting signals the incongruence length difference (ILD) test (
The Akaike information criterion (
The saturation analysis showed insignificant levels of saturation for all three genes, CO1: Iss (0.4398) < Iss.c (0.7384), P = 0.000; 16S: Iss (0.6502) < Iss.c (0.7087), P = 0.007; H3: Iss (0.5591) < Iss.c (0.7193), P = 0.000. The ILD test showed NS conflicting signals between the genes combined: CO1 vs. H3 (P = 0.99), and 16S vs. H3 (P = 0.08), except CO1 vs. 16S (P= 0.001). MRMODELTEST 2.3 selected the models GTR+I+G for CO1 and 16S and GTR+I for H3 (CO1 γ shape = 0.34, proportion of invariant sites = 0.26; 16S γ shape = 0.84, proportion of invariant sites = 0.24; H3 proportion of invariant sites = 0.81).
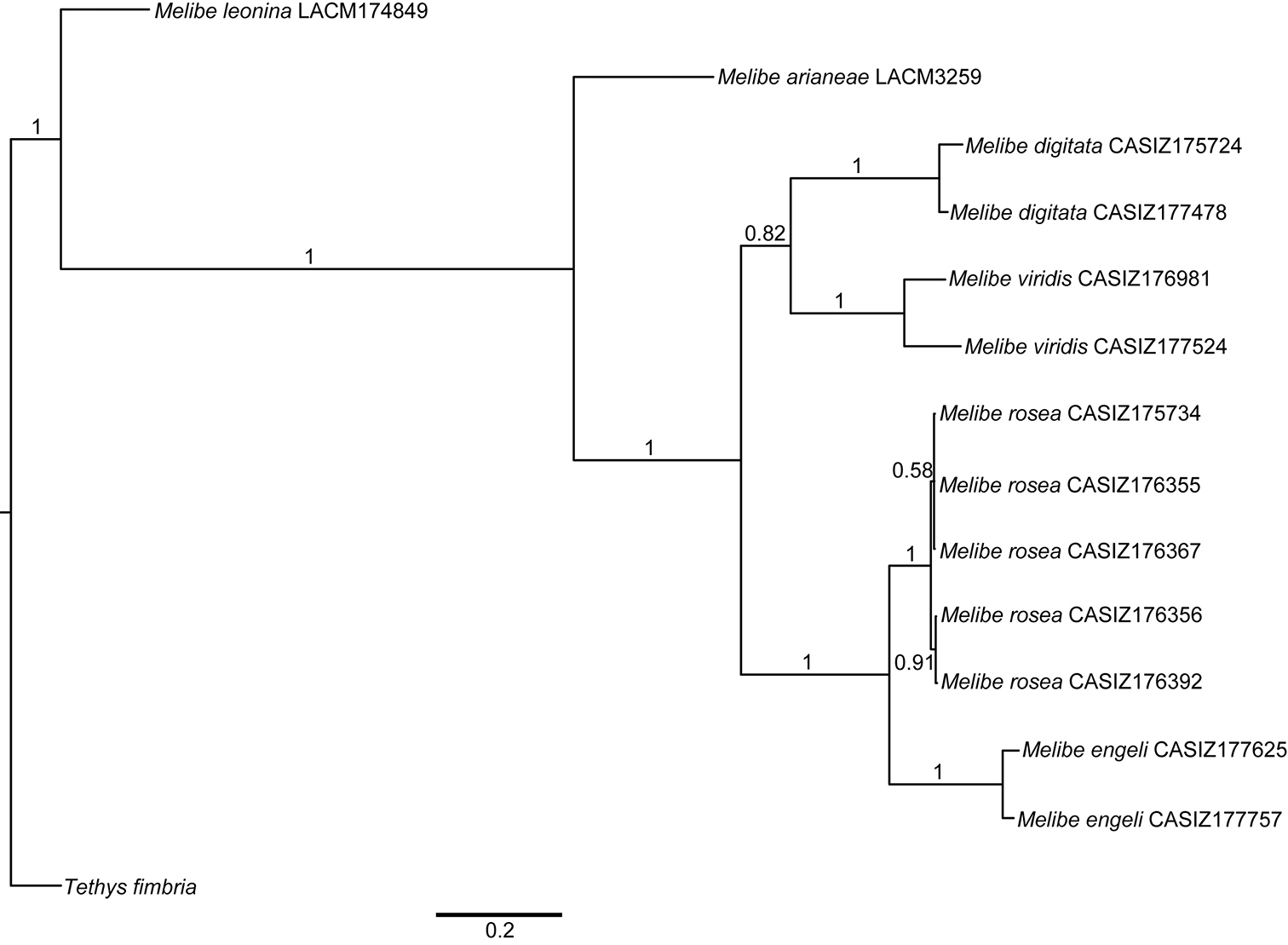
The combined analysis of the three genes (H3, 16S, and CO1) produced a consensus Bayesian tree in which the monophyly of Melibe is well supported, posterior probability (pp) = 1 (Fig. 1). Within Melibe, the temperate species Melibe leonina is the most basal, however this result must be taken cautiously as several other species were not included in the analysis. For the rest of the species analyzed, Melibe arianeae sp. n. is sister to the tropical Indo-Pacific and South African taxa (pp = 1), which includes the Mediterranean non-native Melibe viridis as well as Melibe engeli, the species morphologically more similar to Melibe arianeae. All the species with more than one specimen included in the analysis (Melibe digitata, Melibe viridis, Melibe rosea and Melibe engeli) are monophyletic and well supported (pp = 1).
When aligned with other species of opisthobranchs including Tethys, the CO1 sequence of Melibe arianeae as well as those of other tropical species of Melibe available in GenBank show 4 codon deletions. These codons are in positions 87-89, 352-354, 470-472, and 473-475 (2) of the partial sequence alignment.
Bayesian consensus tree of Melibe including museum voucher numbers and posterior probabilities. Abbreviations: CASIZ, California Academy of Sciences, Invertebrate Zoology; LACM, Natural History Museum of Los Angeles County.
urn:lsid:zoobank.org:act:B9B242B1-9440-4AC3-88D9-A7260986172E
http://species-id.net/wiki/Melibe_arianeae
Figs 2–4Lake Worth Lagoon, Palm Beach County, Florida (26.782781, -80.04468), 3 m depth, 6 mm preserved length, April 6, 2013, A. Dimitris leg.
Holotype preserved in ethanol 95%, dissected but no organs removed (LACM 3258). Paratype preserved in ethanol 95%, dissected, reproductive system in the same vial, stomach on a SEM stub (LACM 3259).
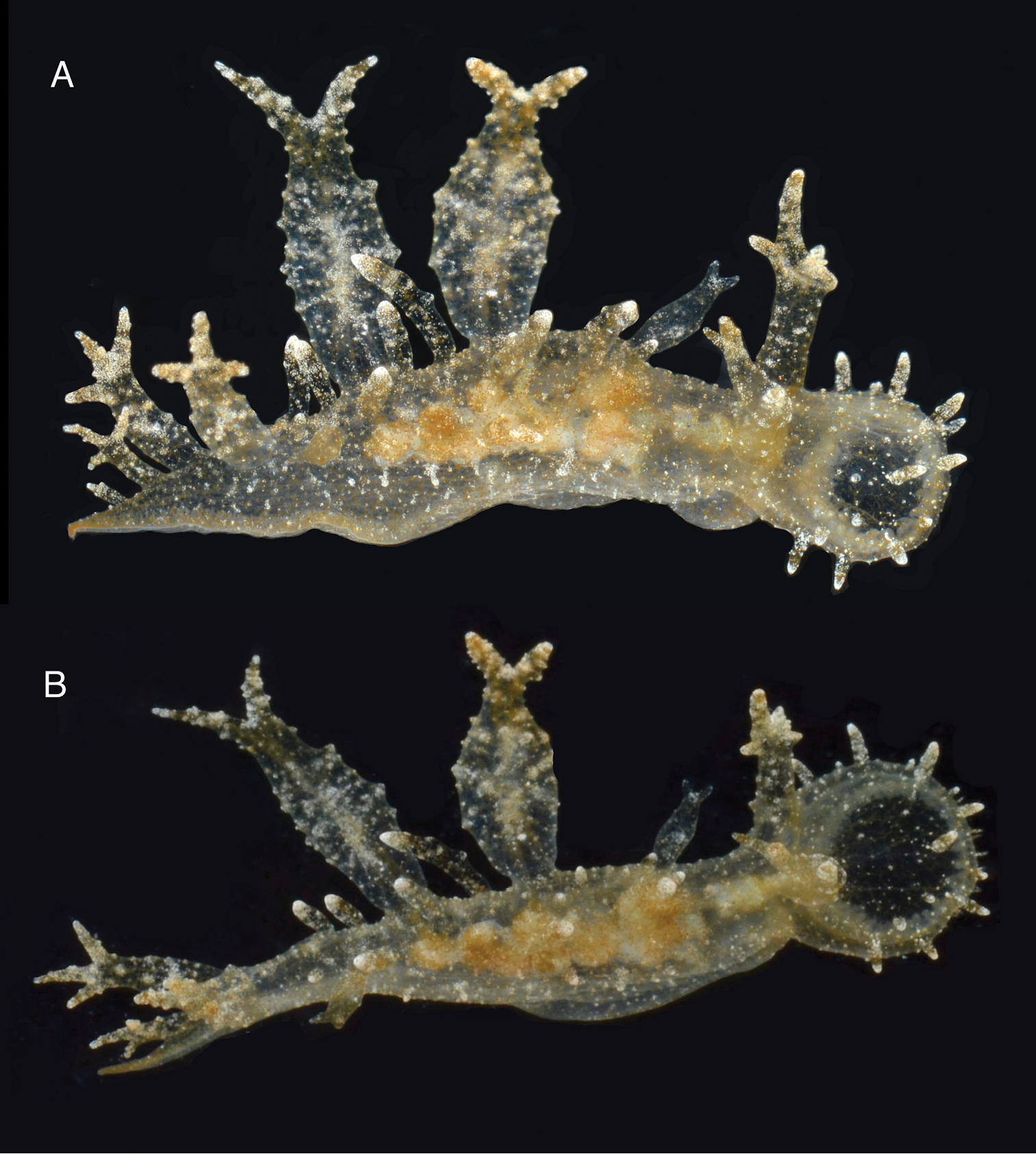
The living animals are nearly transparent, with numerous orange flecks and opaque white blotches all over its surface, and orange-brownish colored internal organs (Fig. 2). The body is limaciform and elongate, somewhat compressed anterolaterally, tapering posteriorly into a long, conical posterior end of the foot. The entire body surface, including cerata and rhinophoral sheaths are covered by numerous minute, opaque white tubercles. In the center of the dorsum of the holotype there are several (8) transparent tentacular papillae of different sizes, also covered with small white tubercles and having opaque white apices. The foot sole is wider anteriorly, it is covered with orange flecks and opaque white blotches as the dorsal surface, but it also has a faint white rim. The circular oral hood is small compared to the rest of the body. The margin of the hood is entire (with no indentations) and bears two rows of elongate papillae. There are papillae on the dorsal surface of the hood, generally resembling those on the body surface, and more concentrated towards the anterior margin. The rhinophores emerge from the posterior end of the oral hood. The rhinophores have 3–4 perfoliations.The rhinophoral sheaths are somewhat inflated and cylindrical, lacking a leaf-like posterior process. The sheaths have 2–3 posterior papillae. The cerata are inflated, oval, completely covered with small tubercles that give it a broadly warty look. Their distal ends of the cerata are either simple, bifurcate or trifurcate, independently of their size. The cerata are transparent, and the branches of the digestive gland within them are visible as a brownish axis. There are seven cerata alternating on each side of the dorsal midline of the holotype. The anus is located dorsol-lateraly, midway between the first and second anterior cerata. The position of the nephroproct could not be determined. The gonopore is lateral, anterior to the anteriormost right ceras. There are no papillae associated with the gonopore.
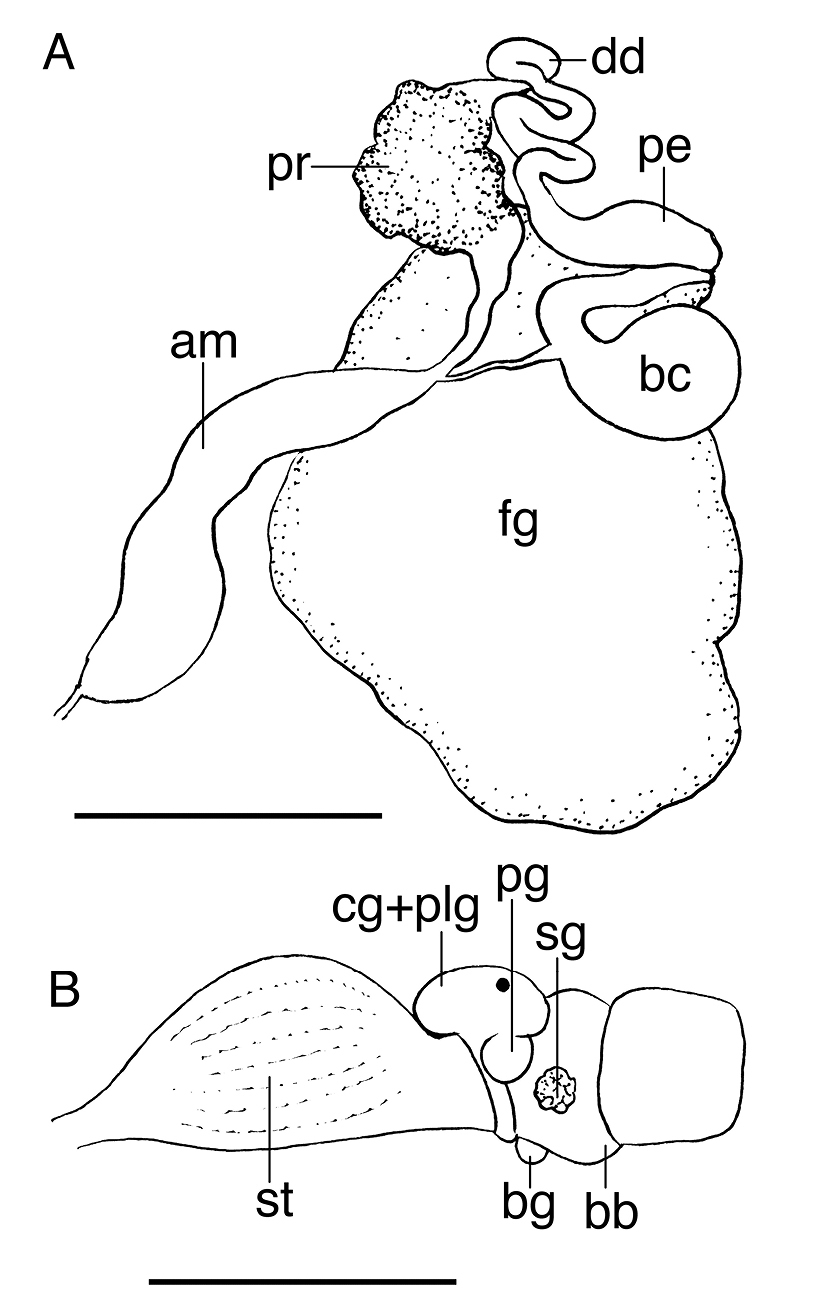
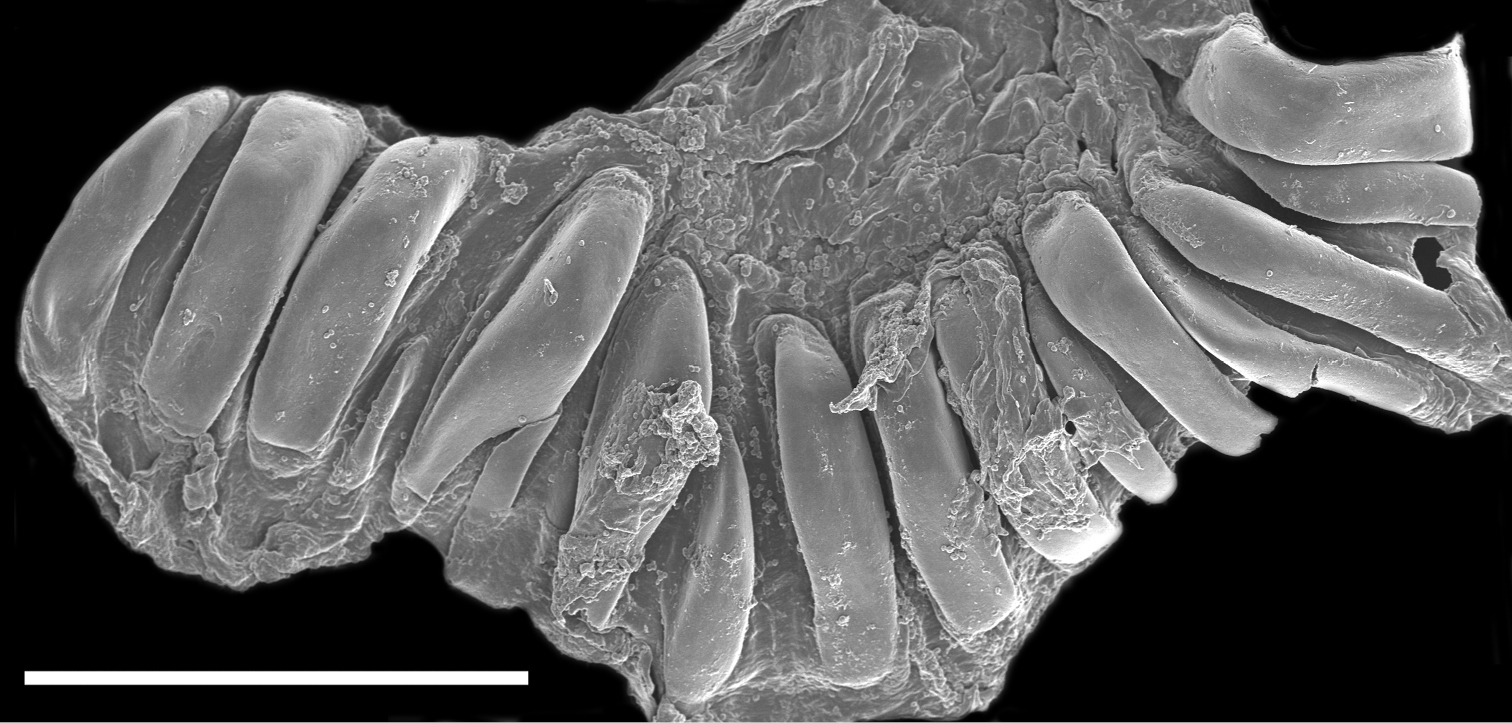
The buccal mass is devoid of a radula but contains a pair of simple, chitinous jaws. The jaws (not illustrated) have smooth borders and lack denticles on the masticatory border. The short esophagus emerges from the posterior end of the buccal mass and expands into a muscular stomach (Fig. 3B). Two small salivary glands are located laterally, one on each side of the buccal mass. The posterior portion of the stomach contains 18 elongate, thick and robust chitinous plates of various sizes (Fig. 4). The reproductive system is triaulic and contains a series of four spherical, well-separated ovotestis bodies, connected to a large ampulla. The ampulla connects into the female gland complex (Fig. 3A) near the point where the prostate emerges. The prostate is a short and wide glandular structure connected to a long, and convoluted deferent duct that expands distally into the penial sac. The vagina is short and wide and connects directly into a large bursa copulatrix. The narrow and straight uterine duct connects to the female gland complex. A serial seminal receptacle (present in other species of Melibe) was not observed. The central nervous system (Fig. 3B) is located above the esophagus and contains a fused pair of cerebral and pleural ganglia, as well as a pair of pedal ganglia. The buccal ganglia are located at the proximal end of the buccal mass.
Two views of the holotype of Melibe arianeae sp. n. (LACM 3258). A. Dorsolateral view showing the right side of the animal. B. Dorsal view showing the oral hood border through the semi-transparent skin.
A Reproductive system of the paratype of Melibe arianeae sp. n. (LACM 3259) B Anterior portion of the digestive system of the holotype of Melibe arianeae sp. n. (LACM 3258). Scale bars = 1mm. Abbreviations: am, ampulla; bb, buccal bulb; bc, bursa copulatrix; bg, buccal ganglion; cg, cerebral ganglion; dd, deferent duct; fg, female gland complex; pe, penis; plg, pleural ganglion; pg, pedal ganglion; pr, prostate; sg, salivary gland; st, stomach.
Scanning electron micrograph of the dissected stomach of the paratype of Melibe arianeae sp. n. (LACM 3259) showing the stomach plates. Scale bar = 500 µm.
This species is named for Ariane Dimitris, amateur naturalist and passionate sea slug enthusiast, who collected the specimens here examined.
Melibe arianeae is well differentiated morphologically relative to other congeners (discussed below). Molecular evidence is provided to support the placement of this new species in Melibe and to compare it to other species for which sequence data are available. There are large gaps in the molecular coverage of Melibe, thus comparison to all other species is not possible at this point. However, the phylogenetic tree here presented shows that Melibe arianeae is sister to a well-supported clade containing some of the most morphologically similar species to Melibe arianeae.
Melibe arianeae is externally most similar to Melibe engeli, originally described from New Caledonia and recently reported from Japan, the Hawaiian Islands and the Philippines (
All other species of Melibe examined and reviewed by
The specimen of Melibe sp. illustrated by
One of the most intriguing aspects of the genetics of Melibe is the presence of four deletions in the sequence of the cytochrome c oxidase subunit 1protein in tropical congeners and Melibe rosea from South Africa. A protein alignment revealed that these deletions resulted in the loss of 4 residues in the cytochrome c oxidase subunit 1 protein. The structure of the cytochrome c oxidase of Paracoccus denitrificans was reconstructed by
Molecular and morphological evidence confirmed that the specimens from Florida here examined belong to Melibe and therefore this paper is the first confirmed record of this group in the tropical western Atlantic Ocean. Morphological examinations also confirmed that these specimens constitute an undescribed species, which is morphologically distinct from other species of Melibe described to date. Additionally, these specimens are genetically distinct from other species of Melibe so far sequenced. Our knowledge of the phylogeny of Melibe is spotty, as many species have not been sequenced yet, thus few conclusions can be drawn from the Bayesian consensus tree. However, it clear that the tropical Indo-Pacific species studied so far form a monophyletic group. The presence of four deletions in the sequence of the cytochrome c oxidase subunit 1 protein in some Melibe could have important implications to understand protein function and selection on mitochondrial genes.
We are extremely grateful to Ariane Dimitris for providing specimens that were essential for the completion this project. Lindsey Groves from the LACM assisted with access to the collection and curation of specimens. This paper was supported by a Cal Poly Pomona Research, Scholarship, and Creative Activity grant and a FuTURE Kellogg Minigrant to A. Valdés as well as a scholarship to E. Espinoza from the NIH MBRS Research Initiative for Scientific Enhancement (RISE) grant to Cal Poly Pomona (2 R25 GM061190-05A2). The SEM work was conducted at the Natural History Museum of Los Angeles County, supported by the National Science Foundation under MRI grant DBI-0216506 to A. Valdés et al., with the assistance of Giar-Ann Kung. Two anonymous reviewers and the editor (Robert Hershler) made constructive comments that substantially improved the manuscript.