(C) 2013 Elkin Y. Suárez-Villota. This is an open access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC-BY), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
The recently described taxon Drymoreomys albimaculatus is endemic to the Brazilian Atlantic Forest and its biology and genetics are still poorly known. Herein, we present, for the first time, the karyotype of the species using classical and molecular cytogenetics, which showed 2n=62, FN=62, and interstitial telomeric signals at the sex chromosomes. Nuclear and mitochondrial DNA sequences from the two karyotyped individuals verify the taxonomic identity as the recently described Drymoreomys albimaculatus and confirm the relationship of the species with other Oryzomyini. Additionally, external morphological information is provided.
Oryzomyini, karyotype, CBG banding, GTG banding, FISH, IRBP, Cyt b
The Atlantic Forest harbors a high diversity of mammals, 20 percent of which are rodents of the subfamily Sigmodontinae (
Recently,
Here, we describe the karyotype of Drymoreomys albimaculatus for the first time. In order to investigate the molecular identification of the two karyotyped animals, we added its Cyt b and IRBP sequences to the molecular data published by
One male and one female were collected with pitfall traps in Santa Virgínia, Parque Estadual da Serra do Mar [45°03.00' to 45°11.00'W (DDM); 23°24.00' to 23°17.00'S (DDM)], state of São Paulo, Brazil. Pelage color and external measurements were taken during the fieldwork. Vouchers of both individuals are deposited in the Coleção de Mamíferos da Universidade Federal do Espírito Santo (UFES) under the catalog numbers UFES 2271 and UFES 2272.
Metaphases were obtained in vivo from spleen and bone marrow, according to
DNA was extracted from liver with Chelex 5% (Bio-Rad) following
We performed maximum likelihood (ML) and Bayesian analyses using concatenated Cyt b-IRBP data set. For both analyses we used gene-specific unlinked models. The best-fitting model of nucleotide substitution for each gene was selected using the Akaike information criterion in accordance with the procedure outlined by
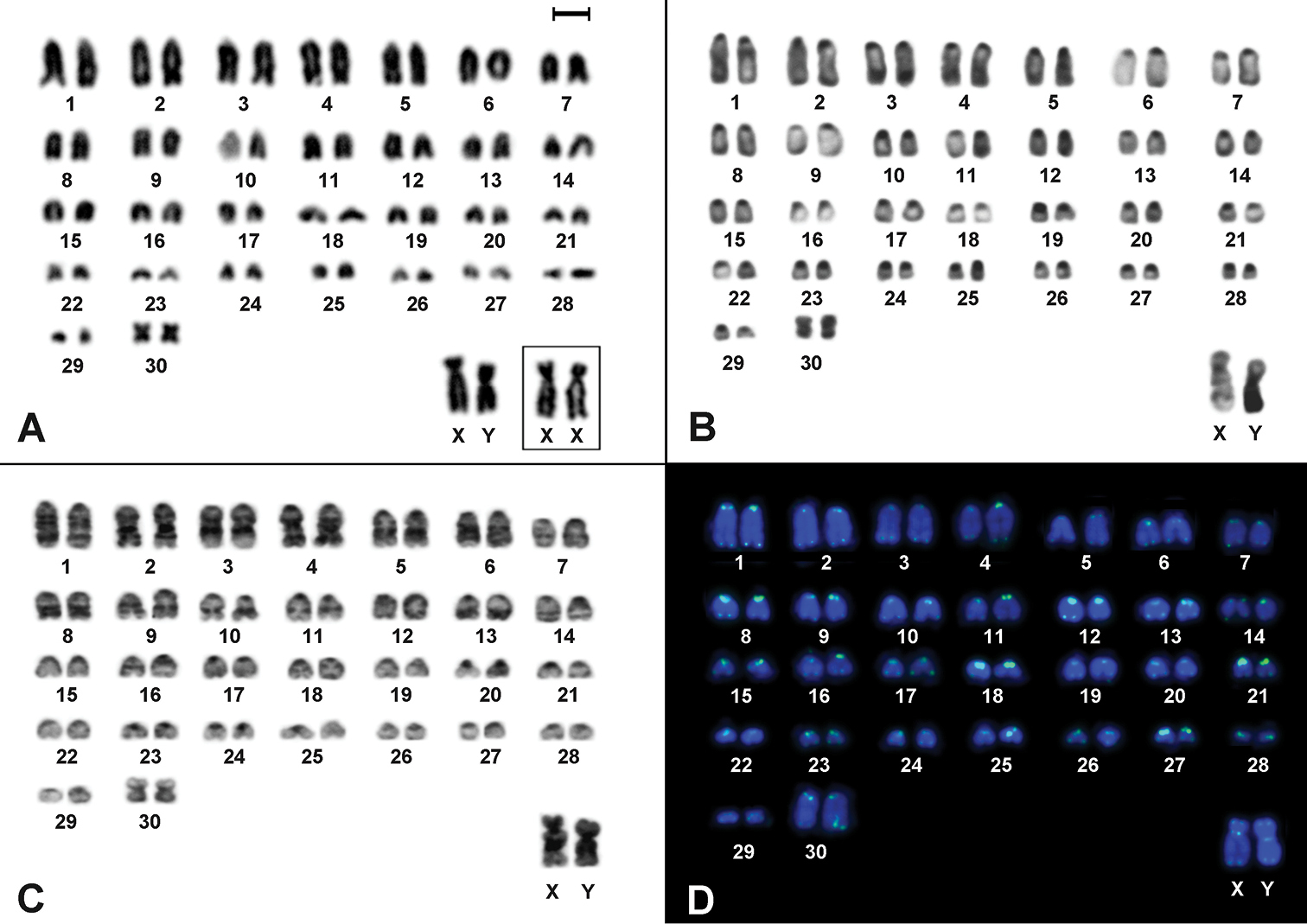
The animals showed 2n=62, FN=62, and the autosome set composed of 29 acrocentric pairs decreasing in size, and one small metacentric pair (Fig. 1A). The X is a large submetacentric, and the Y is a large submetacentric slightly smaller than the X (Fig. 1A). CBG-banding revealed pericentromeric constitutive heterochromatic blocks in all autosomes and in the long arm of Y (Fig. 1B). GTG-banding allowed the identification of almost all autosomic pairs, the X chromosome exhibited two interstitial bands at the long arm while a conspicuous pattern in the Y was not found (Fig. 1C). FISH detected telomeric signals at the ends of all chromosomes and additional telomeric sequences were found in the pericentromeric region of both X and Y chromosomes (Fig. 1D).
Cytogenetic analyses in Drymoreomys albimaculatus from Santa Virgínia, state of São Paulo, Brazil. A Karyotype of male (2n=62, FN=62), after conventional staining. Inset: sex chromosomes of a female B CBG-banding of a male C GTG-banding of a male D Fluorescent in situ hybridization using telomeric PNA probe over male mitotic plates. Bar scale = 10 μm.
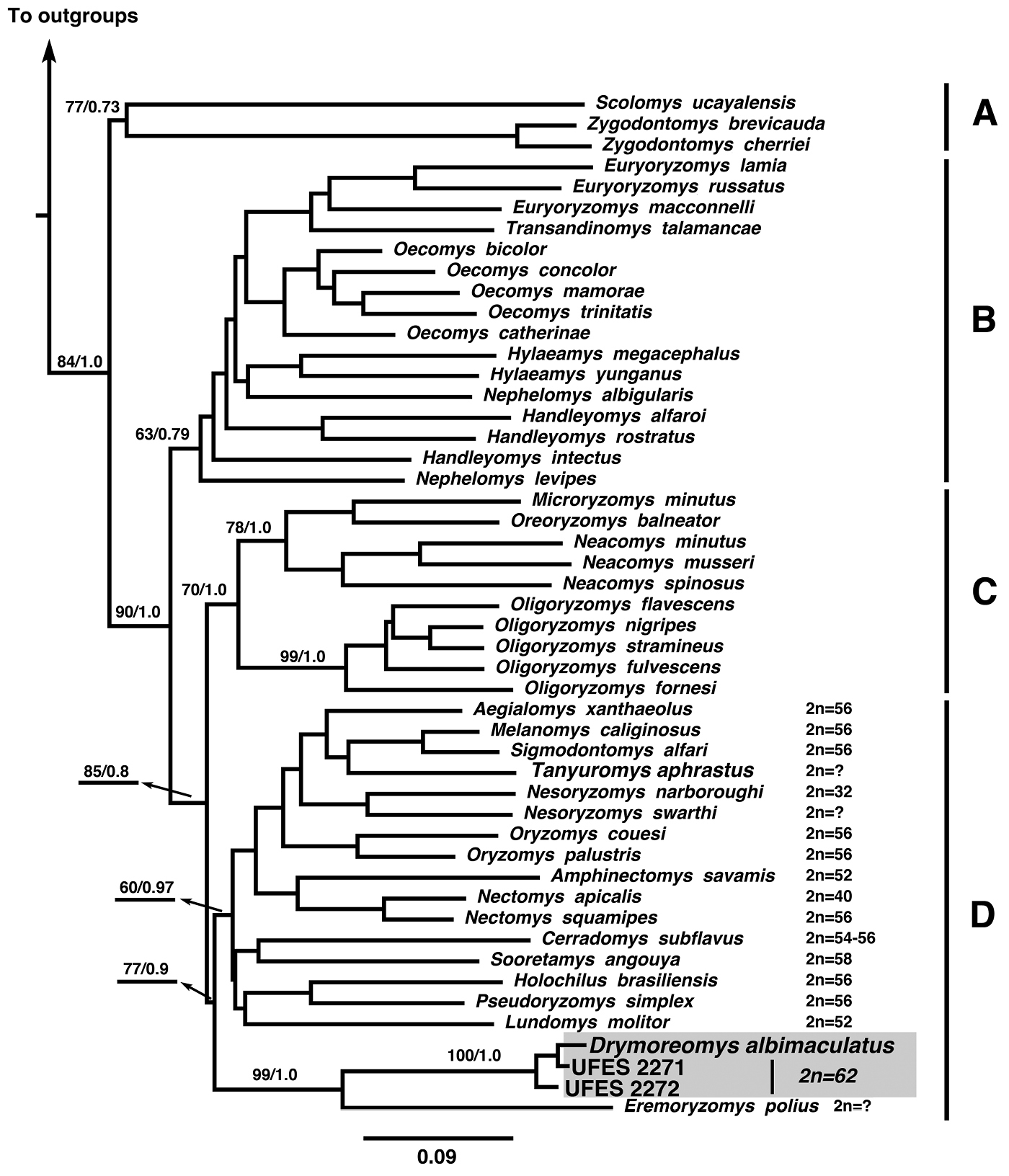
The model selected for the phylogenetic analyses (ML and Bayesian) was GTR +I + Γ for each gene. The best ML tree had a -ln likelihood score of -22, 345.02. The Bayesian analysis recovered a consensus topology similar to the best ML tree and the results recovered the four well-supported clades A, B, C, and D (Fig. 2) previously reported by
Maximum likelihood tree of combined molecular datasets [cytochrome b (Cyt b), interphotoreceptor retinoid binding protein (IRBP)] using Santa Virgínia specimens (UFES2271, UFES2272). Bootstrap nodal support indices and Bayesian posterior probabilities are shown above the branches, respectively. Outgroups include Peromyscus maniculatus (Neotominae); Nyctomys sumichrasti (Tylomyinae), Delomys sublineatus (Sigmodontinae), Thomasomys baeops (Sigmodontinae), and Wiedomys pyrrhorhinos (Sigmodontinae). Available diploid numbers (2n) of clade D are indicated (for details see Table 1), although the lowest diploid number (Nectomys palmipes, 2n= 16,
Phylogenetic analyses (ML and Bayesian) recovered the four clades A, B, C, and D (Fig. 2) recovered by
The diploid number of Drymoreomys albimaculatus corroborates the pattern found for the majority of the Oryzomyini species, in which karyotypes present relatively high chromosome number and predominantly acrocentric pairs. The typical heterochromatic pattern of sex chromosomes is also found in most of the oryzomyine species and it is an essential condition for the recognition of the Y (Fig. 1B).
The karyotype herein reported for Drymoreomys albimaculatus is species-specific, since only three other Oryzomyini species present the same diploid number, but different FN: Oligoryzomys fornesi (2n=62, FN=64), Oligoryzomys delicatus (2n=62, FN=74 and 76), and Oligoryzomys nigripes (2n=62, FN=80, 81 and 82) (
A compilation of karyological studies in representative species of clade D is presented in Table 1. Notably, cytogenetic data in Oryzomyini has increased considerably in the last decades, mainly because the karyotype has become a valid tool for identifying species of this group. Although several species still remain without karyotypic information (e.g., Eremoryzomys polius, Tanyuromys aphrastus, Nesoryzomys swarthy), the diploid number within clade D varies from 16 in Nectomys palmipes (
Species of clade D present sex and supernumerary chromosomes easily identifiable with classical cytogenetic approaches, and some species exhibit sex chromosomes with polymorphisms/heteromorphisms and interstitial telomeric signals (ITS; Table 1, Fig. 1D). Thus, this clade is an excellent model to study origin, evolution, and chromatin composition of these chromosomes. For instance, a superficial morphological comparison among sex chromosomes from Table 1 could suggest the occurrence of pericentric inversions, or whole-arm heterochromatin additions or deletions.
Cytogenetic characteristics of Oryzomyini species of clade D, with diploid number (2n), fundamental number (FN), morphologies of autosomal pairs and sex chromosomes, polymorphisms described and references. *Supernumerary chromosomes are not included in autosomal morphologies. A=acrocentric; M=metacentric; SM=submetacentric; ST=subtelocentric; ITS=interstitial telomeric signals; NA=not available.
| Species | 2n | FN | Autosomal morphologies |
Sex chromosome morphologies | Cytogenetic characteristics | References |
|---|---|---|---|---|---|---|
| Aegialomys xanthaeolus | 56 | 58 | 25 A 2 M/SM |
X: large A Y: small A |
||
| Melanomys caliginosus | 56 | 58 | 25 A 2 M |
X: large ST Y: medium ST |
||
| Sigmodontomys alfari | 56 | 54 | 27 A | X: large A Y: small A |
||
| Tanyuromys aphrastus | NA | NA | NA | NA | ||
| Nesoryzomys narboroughi | 32 | 50 | 5 A 8 M/SM 2 ST |
X: medium A Y: small A |
||
| Nesoryzomys swarthi | NA | NA | NA | NA | ||
| Oryzomys couesi | 56 | 56 | 26 A 1 M |
X: large SM Y: medium A/ST |
Y heteromorphisms | |
| Oryzomys palustris | 56 | 56 | 26 A 1 M |
X: large A Y: minute A |
||
| Amphinectomys savamis | 52 | 66 | NA | NA | ||
| Nectomys apicalis | 42 | 40 | 20 A | X and Y: A | ||
| Nectomys squamipes | 56–59 | 56 | 26 A 1 M |
X: large SM/ ST Y: medium/ small SM/ST |
0-2 B chromosomes; sex chromosomes polymorphisms | |
| Cerradomys subflavus | 54–56 | 62 | 21 A, 3 SM, 2M 23 A, 2 SM, 2M |
X: large A/ ST Y: medium A/ large A |
Centric fusion/fission, pericentric inversion, sex chromosomes polymorphisms | |
| Sooretamys angouya | 58, 60 | 60, 64 | 26 A 2 M |
X: large A Y: medium ST |
0 or 2 B chromosomes | |
| Holochilus brasiliensis | 56–58 | 56, 58, 60 | 26 A 1 M |
X: large ST Y: small SM |
0 to 2 B chromosomes | |
| Pseudoryzomys simplex | 56 | 54, 55 | 27 A | X: large A Y: medium A |
Heteromorphic pair 17 due to addition of constitutive heterochromatin | |
| Lundomys molitor | 52 | 58 | 21 A 4 M |
X: large SM Y: small M |
X heteromorphism | |
| Drymoreomys albimaculatus | 62 | 62 | 29 A 1M |
X: large SM Y: medium SM |
ITS in both sex chromosomes | Present study |
| Eremoryzomys polius | NA | NA | NA | NA | NA |
The specimens collected were medium sized (male body mass: 46.5 g, head and body length: 115 mm, and tail length: 142 mm; female body mass: 57 g, head and body length: 127 mm, and tail length: 170 mm). Tail was longer than head and body, and was a uniform color on both sides. Male hind footwas short (25 mm, 22% of head and body length) and ears were small (16 mm; 14% of head and body length). These external morphological measures overlapped with those of the Drymoreomys albimaculatus holotype (
This work was supported by FAPESP (2008/00493-9 and 2010/03432-0 for CDN, Jovem Pesquisador 2005/04557-3 for MJJS), CNPq (131891/2008-1 for CLN) and by Conicyt (Becas Chile 74130017 for EYSV). We thank two anonymous reviewers for their helpful comments.