(C) 2013 Karen Cox. This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
Citation: Cox K, Thomaes A, Antonini G, Zilioli M, De Gelas K, Harvey D, Solano E, Audisio P, McKeown N, Shaw P, Minetti R, Bartolozzi L, Mergeay J (2013) Testing the performance of a fragment of the COI gene to identify western Palaearctic stag beetle species (Coleoptera, Lucanidae). In: Nagy ZT, Backeljau T, De Meyer M, Jordaens K (Eds) DNA barcoding: a practical tool for fundamental and applied biodiversity research. ZooKeys 365: 105–126. doi: 10.3897/zookeys.365.5526
The taxonomy of stag beetles (Coleoptera: Lucanidae) remains challenging, mainly due to the sexual dimorphism and the strong allometry in males. Such conjecture confounds taxonomic based conservation efforts that are urgently needed due to numerous threats to stag beetle biodiversity. Molecular tools could help solve the problem of identification of the different recognized taxa in the “Lucanus cervus complex” and in some related Palaearctic species. We investigated the potential use of a 670 bp region at the 3’ end of the mitochondrial cytochrome c oxidase subunit I gene (COI) for barcoding purposes (different from the standard COI barcoding region). Well resolved species and subspecies were L. tetraodon, L. cervusakbesianus, L. c. laticornis, as well as the two eastern Asian outgroup taxa L. formosanus and L. hermani. Conversely, certain taxa could not be distinguished from each other based on K2P-distances and tree topologies: L. c. fabiani / L. (P.) barbarossa, L. c. judaicus / an unknown Lucanus species, L. c. cervus / L. c. turcicus / L. c. pentaphyllus / L. (P.) macrophyllus / L. ibericus. The relative roles of phenotypic plasticity, recurrent hybridisation and incomplete lineage sorting underlying taxonomic and phylogenetic discordances are discussed.
Lucanus spp., Stag beetle, Western Palaearctic, DNA barcoding, COI
Lucanidae Latreille, 1804 is a family of Coleoptera showing in most species pronounced sexual dimorphism and strong external morphological allometry in males. The species of the Holarctic and Oriental distributed genus Lucanus Scopoli, 1763 are renowned for the striking appearance of the males. With their large body size and prominent mandibles, the male stag beetles are very popular among amateur entomologists and as terrarium pets, mainly in Japan. Currently, there are more than 90 Lucanus species described, however, validity of these designations is considered questionable in many cases. Sexual dimorphism and size variation complicate the taxonomy (
The genus Lucanus is subdivided into the subgenera Lucanus sensu stricto and Pseudolucanus Hope & Westwood, 1845. Members of the latter have a peculiar stout body and substantial analogy of morphology that makes it quite easy to distinguish them from members of the subgenus Lucanus (
The distribution of many of these taxa remains poorly resolved, however, we can consider some of them as endangered. The practice of removing old trees and dead wood in past and current forest management, has had detrimental effects on this group of saproxylic beetles (
More detailed information on the distribution and ecology of this species group is needed to get a clear view on their conservation status. But unless the problem of identification of European and West Asian Lucanus is solved, it becomes difficult to set specific conservation priorities, without which rare, neglected and endangered species or Evolutionarily Significant Units (ESUs) may be unrecognised and thus, not given adequate conservation prioritisation (
Lucanus cervus has the widest geographical distribution in the genus and is very variable in form, size and colour (
Lucanus ibericus can be found from Albania to Iran and is sometimes considered a synonym of Lucanus orientalis. Unlike Lucanus cervus, Lucanus ibericus is entirely reddish brown, has a pronotum without a smooth discal line, but with a sinuate posterior and distinct toothed posterior angles (non-sinuate pronotum and blunt angles in Lucanus cervus). The mandibles of the males, which are shorter than those of a typical male Lucanus cervus of equal size, can have an apex with two equal teeth or with the inner tooth fainted and a large internal denticle in the middle. In addition, Lucanus ibericus has six, rarely five, long lamellae on the antennal club.
Lucanus tetraodon described from France, Italy, North Africa, Albania and Greece, can be perceived as a central Mediterranean species. In contrast to Lucanus cervus and Lucanus ibericus, the basal denticle of the mandibles of Lucanus tetraodon is placed in the lower half. Like Lucanus ibericus, the pronotal sides have sharp posterior angles, but the pronotal disc misses the central smooth line. Lucanus tetraodon has six, occasionally five, lamellae on the antennal club. Lucanus tetraodon is by some authors subdivided in subspecies Lucanus tetraodon argeliensis Maes, 1995 in North Africa, Lucanus tetraodon provincialis Colas, 1949 in South France, Lucanus tetraodon corsicus Gautier des Cottes, 1860 in Corsica, Lucanus tetraodon sicilianus Planet, 1899 in Sicily and finally Lucanus tetraodon tetraodon Thunberg, 1806 elsewhere. In addition, specimens of problematic populations of Lucanus cervus from a series of localities in central Italy (northern Latium and Umbria), are known to exhibit apparently intermediate morphological characters between Lucanus cervus and Lucanus tetraodon, which are sympatric in these areas (
The Pseudolucanus species all have six long lamellae forming the antennal club, their body is stout and entirely reddish or blackish brown. Included in this study are Lucanus (Pseudolucanus) barbarossa from the Iberian peninsula and the Maghreb, and Lucanus (Pseudolucanus) macrophyllus reported in south-west Turkey.
A large number of entomologists was contacted to obtain material from the different taxa and from different regions. The samples included whole beetles, especially in regions where identification is problematic, as well as parts of a beetle, sometimes found as road kill or as prey leftovers from birds. Samples were dried and kept at room temperature or preserved in absolute ethanol. In total 76 samples were collected. The species identification was performed, using comparative material and available identification keys. Six samples from Israel and Lebanon could not be identified to species. These unidentified Lucanus specimens have a shape resembling in general the medium to small males of Lucanus cervus akbesianus but with a mandibular structure similar to that of Lucanus cervus turcicus (Zilioli et al. unpublished data). The tissue samples used for DNA extraction depended on what was available, but were mostly legs, which contain large muscles and are therefore rich in mitochondrial DNA (mtDNA). DNA was extracted from ground samples with the E.Z.N.A.® Forensic DNA Kit (Omega Bio-Tek), except for samples K1 and U6 (Table 1) from which DNA was extracted following the salting out procedure described by
List of samples included in the analysis. Primers used are denoted with ‘1’: C1-J-2183 and TL2-N-3014; ‘2’: LCint1F, LCint2F, LCint3F and LCint4F (for sample SB6 also the reverse primers were used); ‘3’: F - 5’ GGGGCATCAGTAGACCTAGC 3’ and R – 5’ TTCAGCAGGTGGTATTAGTTGG 3’.
| Species / subspecies | Code | Primers | Haplotype | GenBank acc. no. | Country | Latitude | Longitude | Date of sampling | Type of conservation | Gender |
|---|---|---|---|---|---|---|---|---|---|---|
| Lucanus cervus akbesianus | UA1 | 1 | UA1 | KF737127 | Turkey | 37.721833, 30.828278 | Jun 2010 | ethanol | Female | |
| UA2 | 1 | UA2 | KF737128 | Turkey | 37.721833, 30.828278 | Jun 2010 | ethanol | Male | ||
| UA3 | 1 | UA3 | KF737129 | Turkey | 37.721833, 30.828278 | Jun 2010 | ethanol | Male | ||
| UA4 | 1 | UA4 | KF737130 | Turkey | 37.676200, 35.862100 | 2010 | ethanol | Male | ||
| UA5 | 1 | UA5 | KF737131 | Turkey | 37.676200, 35.862100 | 2010 | ethanol | Male | ||
| UX1 | 2 | UX1 | KF737132 | Turkey | 36.900000, 31.000000 | Jun 2010 | ethanol | Male | ||
| U10 | 1 | U10 | KF737125 | Turkey | 37.721833, 30.828278 | Jun 2010 | ethanol | Male | ||
| U11 | 1 | U10 | KF737126 | Turkey | 37.721833, 30.828278 | Jun 2010 | ethanol | Male | ||
| Lucanus cervus cervus | A1 | 1 | A1 | KF737071 | Belgium | 50.772652, 4.537656 | Jul 2008 | ethanol | Male | |
| A3 | 1 | A3 | KF737072 | Belgium | 50.736622, 4.331784 | Jun 2009 | ethanol | Female | ||
| C1 | 2 | C1 | KF737093 | Czech rep. | 48.797935, 16.803576 | May 2009 | ethanol | Male | ||
| D13 | 2 | A3 | KF737078 | France | 45.391800, 1.139310 | Jul 2010 | ethanol | Male | ||
| D4 | 1 | D4 | KF737088 | France | 43.458090, 1.431787 | Aug 2010 | ethanol | Male | ||
| D22 | 1 | D22 | KF737092 | France | 47.861145, 2.820327 | 2009 | ethanol | Female | ||
| F12 | 1 | A3 | KF737079 | Greece | 39.808333, 22.653889 | Jun 2009 | ethanol | Female | ||
| F16 | 1 | F16 | KF737083 | Greece | 39.808333, 22.653889 | Jun 2009 | ethanol | Female | ||
| F23 | 1 | F23 | KF737082 | Greece | 39.762333, 21.663281 | Jun 2009 | ethanol | Male | ||
| G3 | 2 | G3 | KF737081 | Hungary | 47.701586, 18.834592 | Jul 2009 | ethanol | Female | ||
| I2 | 1 | I2 | KF737084 | Italy | 45.779241, 8.732981 | Jun 2009 | ethanol | Male | ||
| I3 | 1 | A3 | KF737080 | Italy | 45.779241, 8.732981 | Jun 2009 | ethanol | Male | ||
| I4 | 1 | I4 | KF737085 | Italy | 45.779241, 8.732981 | Jun 2009 | ethanol | Male | ||
| N3 | 1 | N3 | KF737086 | Portugal | 38.795900, -9.397390 | Jul 2010 | ethanol | Male | ||
| O9 | 3 | O9 | KF737087 | Romania | 47.102400, 24.450700 | |||||
| S15 | 1 | S15 | KF737094 | Spain | 40.385100, -6.608460 | Aug 2009 | ethanol | Male | ||
| S19 | 1 | A3 | KF737076 | Spain | 43.304009, -4.814970 | Jul 2009 | ethanol | Female | ||
| V2 | 1 | A3 | KF737077 | UK | 52.028936, 1.067369 | Aug 2009 | dried | Female | ||
| V26 | 3 | V26 | KF737091 | UK | 50.966300, -0.209294 | |||||
| V44 | 3 | V44 | KF737089 | UK | 51.260100, 0.844280 | |||||
| W9 | 2 | W9 | KF737090 | Ukraine | 49.826900, 36.325800 | Jun 2007 | dried | Male | ||
| X1 | X1 | FJ606555 | France | ( |
||||||
| Lucanus cervus fabiani | D11 | 1 | D11 | KF737121 | France | 43.195300, 5.753740 | Jun 2010 | ethanol | Male | |
| Lucanus cervus judaicus | UJ1 | 1 | UJ1 | KF737112 | Turkey | 37.068100, 36.261600 | Jul 2010 | dried | Male | |
| Lucanus cervus laticornis | UL2 | 1 | UL2 | KF737119 | Turkey | 36.875669, 30.457431 | Jun 2007 | ethanol | Male | |
| UL3 | 1 | UL3 | KF737120 | Turkey | 37.763600, 30.558900 | 1995 | dried | Male | ||
| Lucanus cervus pentaphyllus | C2 | 1 | A3 | KF737075 | Czech rep. | 48.797935, 16.803576 | May 2009 | ethanol | Male | |
| F13 | 1 | F13 | KF737104 | Greece | 39.808333, 22.653889 | Jun 2009 | ethanol | Female | ||
| I1 | 1 | A3 | KF737073 | Italy | 45.779241, 8.732981 | Jun 2009 | ethanol | Male | ||
| W7 | 2 | A3 | KF737074 | Ukraine | 48.950200, 38.497600 | Jul 2002 | dried | Male | ||
| Lucanus cervus turcicus | B1 | 1 | B1 | KF737096 | Bulgaria | 42.162733, 27.737650 | Jul 2009 | ethanol | Male | |
| B2 | 1 | B2 | KF737098 | Bulgaria | 41.407800, 25.578583 | Jul 2009 | ethanol | Male | ||
| B7 | 1 | B7 | KF737099 | Bulgaria | 42.060792, 27.977000 | Jul 2009 | ethanol | Male | ||
| B9 | 1 | B1 | KF737097 | Bulgaria | 42.120183, 27.900405 | |||||
| F15 | 2 | F15 | KF737105 | Greece | 39.808333, 22.653889 | Jun 2009 | ethanol | Male | ||
| F7 | 1 | F7 | KF737107 | Greece | 39.866667, 22.733333 | Jun 2009 | ethanol | |||
| F8 | 2 | F7 | KF737108 | Greece | 39.866667, 22.733333 | Jun 2009 | ethanol | |||
| F9 | 1 | F9 | KF737106 | Greece | 39.808333, 22.653889 | Jun 2009 | ethanol | Female | ||
| F11 | 1 | F11 | KF737100 | Greece | 39.808333, 22.653889 | Jun 2009 | ethanol | Male | ||
| F17 | 2 | F17 | KF737101 | Greece | 39.808333, 22.653889 | Jun 2009 | ethanol | |||
| F20 | 1 | F20 | KF737102 | Greece | 39.808333, 22.653889 | Jun 2009 | ethanol | Male | ||
| F21 | 1 | F21 | KF737103 | Greece | 39.808333, 22.653889 | Jun 2009 | ethanol | Male | ||
| U3 | 2 | U3 | KF737109 | Turkey | 41.800000, 27.950000 | Jul 2009 | ethanol | Male | ||
| Unknown species of Lucanus | H1 | 2 | H1 | KF737116 | Israel | 33.217100, 35.753500 | Aug 2009 | ethanol (after freezing) | Male | |
| H2 | 1 | H2 | KF737113 | Israel | 33.217100, 35.753500 | Aug 2009 | dried | Female | ||
| H3 | 2 | H3 | KF737117 | Israel | 33.217100, 35.753500 | Jul 2009 | dried | Male | ||
| H4 | 1 | H4 | KF737114 | Israel | 33.217100, 35.753500 | Jul 2009 | dried | Male | ||
| H5 | 2 | H5 | KF737115 | Israel | 32.959600, 35.864500 | 1998 | dried | Male | ||
| J2 |
2 | J2 | KF737118 | Lebanon | Jul 2009 | dried | Male | |||
| Lucanus ibericus | U6 | 1 | U6 | KF737110 | Turkey | 40.290300, 38.424200 | ||||
| Lucanus tetraodon provincialis | D6 | 1 | D6 | KF737111 | France | 43.066700, 5.850000 | Jun 2010 | ethanol | Male | |
| Lucanus tetraodon | X2 | X2 | EF487727 | ( |
||||||
| Lucanus (Pseudolucanus) barbarossa | SB1 | 1 | SB1 | KF737122 | Spain | 40.828139, -3.831811 | Jul 2004 | dried, later on ethanol | Male | |
| SB6 |
2 | SB6 | KF737124 | Spain | 41.067361, -3.585322 | Sep 2010 | ethanol | Female | ||
| SB7 | 1 | SB7 | KF737123 | Spain | 36.885000, -3.982000 | May 2010 | ethanol | Male | ||
| Lucanus (Pseudolucanus) macrophyllus | UB1 |
2 | UB1 | KF737095 | Turkey | 36.501944, 33.089167 | Aug 2006 | dried | Male | |
| Dorcus parallelipipedus | 1 | K1 | KF737133 | Montenegro | ||||||
| X3 | DQ156023 | ( |
||||||||
| Lucanus formosanus | X4 | FJ606632 | ( |
|||||||
| X5 | FJ606630 | |||||||||
| X6 | FJ606628 | |||||||||
| X5 | FJ606626 | |||||||||
| X5 | FJ606624 | |||||||||
| X5 | FJ606622 | |||||||||
| X8 | FJ606583 | |||||||||
| Lucanus hermani | X9 | FJ606552 | ( |
|||||||
† sequences with a maximum of seven double peaks.
We first attempted to sequence the COI barcoding region with the primers developed by Folmer et al. (1994) on a subset of samples. Despite PCR optimization trials, amplification of this fragment largely failed. Instead, a 800 bp fragment of the 3’ end of the COI gene was amplified using the primer set C1-J-2183 (5’ CAACATTTATTTTGATTTTTTGG 3’) and TL2-N-3014 (5’ TCCAATGCACTAATCTGCCATATTA 3’) (
COI sequences available on GenBank were added. The COI sequence of Lucanus cervus cervus obtained by
DNA sequences have been deposited in GenBank under accession numbers KF737071 to KF737133 (Table 1).
Overall quality of the sequences was evaluated manually. Only samples with high quality chromatograms for at least 300 bp were retained for further analyses. Sequences were aligned by hand and using CLUSTALW v1.4 (
As K2P-distance is the most commonly used distance metric in DNA barcoding (
Of a total of 76 samples, thirteen samples with low quality sequences were removed: five Lucanus cervus cervus, one Lucanus cervus pentaphyllus, three Lucanus cervus turcicus and four Lucanus (Pseudolucanus) barbarossa. Three sequences showed a few double peaks: one Lucanus (Pseudolucanus) barbarossa (SB6: 5 ambiguous sites), one Lucanus (Pseudolucanus) macrophyllus (UB1: 7 ambiguous sites) and one unidentified species of Lucanus (J2: 2 ambiguous sites) (Table 1). None exhibited indels or stop codons which are indicative of the presence of NUMTs (
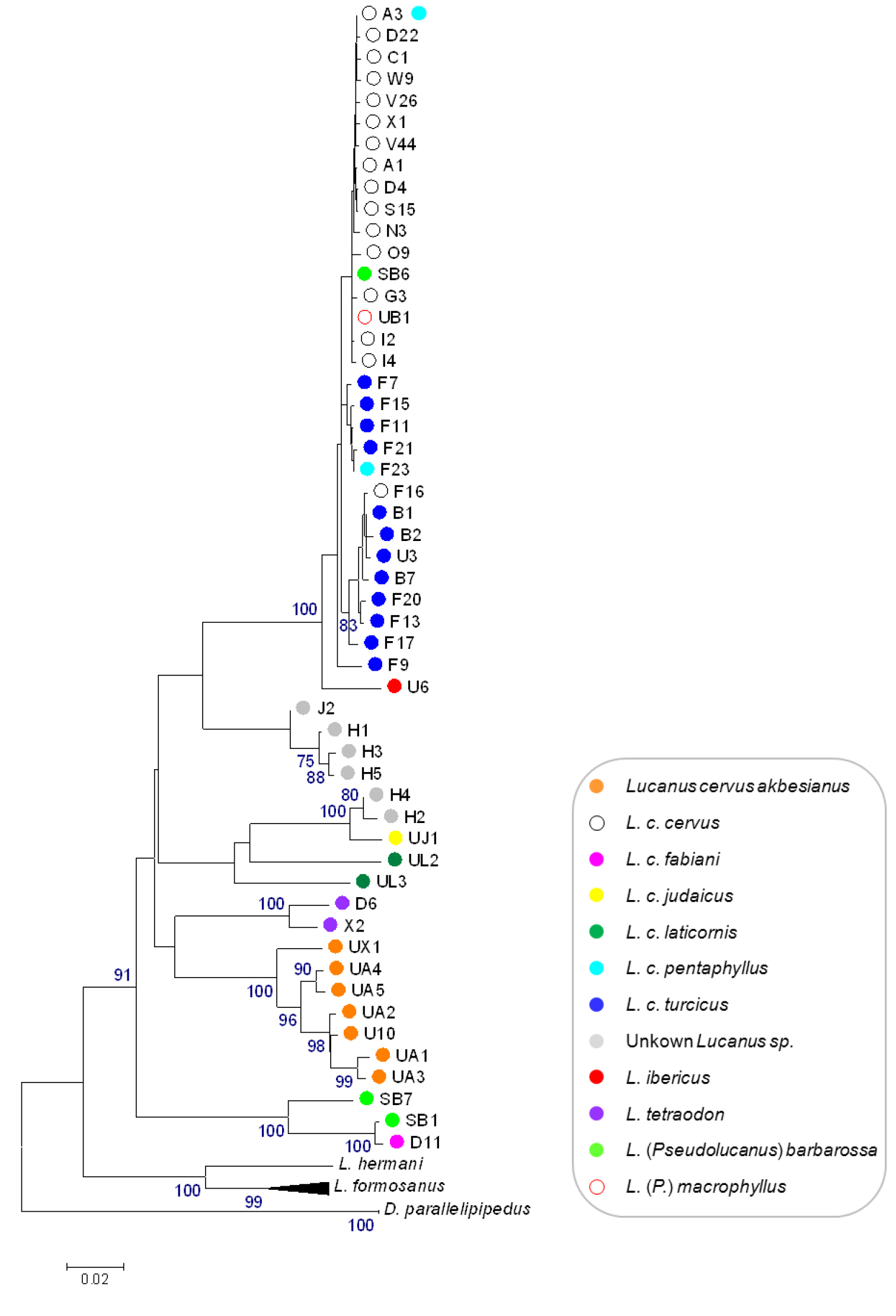
Both the NJ tree and the BI tree showed the same overall configuration (Figure 1 and Appendix 1, respectively) except for the position of the unidentified Lucanus specimens. In the NJ tree these specimens fall into two clusters with unresolved affinities (Figure 1). In the BI tree they form a single well-supported clade together with specimens identified as Lucanus cervus judaicus and Lucanus cervus laticornis (Appendix 1). The unidentified specimens fail to form a single monophyletic cluster as one subclade also includes Lucanus cervus judaicus. The BI tree showed Lucanus cervus laticornis to be monophyletic with probability 1, instead of paraphyletic as was shown in the NJ tree with bootstrap support below 70%. In both trees, several species as well as subspecies fall into distinct clades, whereas Lucanus cervus cervus, Lucanus cervus turcicus, Lucanus cervus pentaphyllus, Lucanus (Pseudolucanus) macrophyllus and Lucanus ibericus cluster in the same shallow clade (called the ‘Lucanus cervus cervus clade’ hereafter). In addition, three out of four samples of Lucanus cervus pentaphyllus share a haplotype with Lucanus cervus cervus (haplotype A3) which is the most common haplotype among Lucanus cervus sequences (Table 1). Within this clade Lucanus cervus cervus, Lucanus cervus turcicus and Lucanus cervus pentaphyllus are polyphyletic. Unexpectedly, one sample of Lucanus (Pseudolucanus) barbarossa and the sample of Lucanus (Pseudolucanus) macrophyllus are also embedded in this clade. Looking at the sequences, they only differ from haplotype A3 at their five and seven ambiguous sites, respectively. Because the two other specimens of Lucanus (Pseudolucanus) barbarossa form a separate clade with Lucanus cervus fabiani, sample SB6 is excluded from further calculations but will be discussed below.
Bootstrap consensus NJ tree inferred from 10 000 replicates, with a cut off value of 70%, based on K2P-distances between 60 haplotypes of the 3’ end of the COI gene.
The nucleotide composition of all the sequences was AT-rich, with 29.5% A, 35.2% T, 15.5% G and 19.7% C. There were 36.4% nucleotide sites variable and 12.1% variable amino acid sites, of which 94.3% and 77.8% were parsimony informative, respectively. When Dorcus parallelipipedus was excluded from the dataset, variable sites decreased to 33.3% for nucleotides and 7.2% for amino acids (94.2% and 56.2% parsimony informative, respectively). Nucleotide composition and K2P-distances calculated for each codon position are shown in Table 3.
Although specimen J2 of the unidentified specimens of Lucanus clustered with the other specimens of the same taxon in the NJ and BI trees, the pairwise interspecific K2P-distances with J2 differed substantially from those with H1 to H5 (comparisons with Lucanus cervus judaicus not included). More specifically, the minimum pairwise interspecific K2P-distance between J2 and the other unidentified taxa was 0.064 opposed to 0.087–0.095 when taking H1 to H5 into account. J2 is one of three incomplete sequences and missing information from position 179 to 399 in the sequence of J2 where several simple nucleotide diagnostics are present (Appendix 2). Therefore, this sample was removed from the dataset for subsequent analysis.
The congeneric interspecific K2P-distances between the western Palaearctic taxa and the eastern Asian species Lucanus formosanus and Lucanus hermani range from 0.156 to 0.198. Distances between taxa of Lucanus and Dorcus went from 0.211 until 0.259. K2P-distances within and between the investigated western Palaearctic taxa of Lucanus are shown in Table 2. As indicated by the NJ and BI trees, the taxa Lucanus cervus cervus, Lucanus cervus pentaphyllus, Lucanus cervus turcicus and Lucanus (Pseudolucanus) macrophyllus cannot be distinguished based on the COI fragment; K2P-distances range from 0 to 0.021, and all taxa are reciprocally polyphyletic. Whereas the first three subspecies of Lucanus cervus are distinguished solely on the basis of the number of lamellae on the antennal club, Lucanus (Pseudolucanus) macrophyllus is morphologically much more distinctive, resembling Lucanus ibericus. Although Lucanus ibericus is part of the Lucanus cervus cervus clade, it shows slightly higher K2P-distances with the other members of this clade (0.028–0.032). Note that we only had a single specimen. Moderate to relatively high within (sub)species distances were found for Lucanus cervus laticornis (0.085), certain specimens of the unidentified Lucanus sp. (max. 0.054) and Lucanus (Pseudolucanus) barbarossa (0.53). On the other hand, between the latter and Lucanus cervus fabiani a small to moderate distance exists (0.004 and 0.058). This is also the case between taxa H2 and H4 of the unknown Lucanus sp. and Lucanus cervus judaicus (K2P-distance of 0.018 and 0.016, respectively). The remaining distances between (sub)species ranged from 0.087 and 0.179.
Intra- and interspecific K2P-distances for the 670 bp COI gene of western Palaearctic Lucanus (sub)species. NA: intraspecific K2P-distance cannot be presented because only one sample is available.
| Lucanus cervus cervus | Lucanus cervus pentaphyllus | Lucanus cervus turcicus | Lucanus cervus fabiani | Lucanus cervus akbesianus | Lucanus cervus judaicus | Lucanus cervus laticornis | Lucanus ibericus | Lucanus tetraodon | Lucanus (Pseudolucanus) macrophyllus | Lucanus (Pseudolucanus) barbarossa | unknown Lucanus sp. | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lucanus cervus cervus | 0–0.018 | |||||||||||
| Lucanus cervus pentaphyllus | 0–0.018 | 0–0.014 | ||||||||||
| Lucanus cervus turcicus | 0.001–0.021 | 0.003–0.017 | 0–0.017 | |||||||||
| Lucanus cervus fabiani | 0.161–0.167 | 0.160–0.163 | 0.159–0.169 | NA | ||||||||
| Lucanus cervus akbesianus | 0.118–0.161 | 0.121–0.155 | 0.121–0.165 | 0.159–0.174 | 0–0.045 | |||||||
| Lucanus cervus judaicus | 0.151–0.164 | 0.153–0.160 | 0.155–0.170 | 0.167 | 0.144–0.154 | NA | ||||||
| Lucanus cervus laticornis | 0.134–0.160 | 0.134–0.155 | 0.132–0.164 | 0.162–0.165 | 0.135–0.150 | 0.089–0.094 | 0.085 | |||||
| Lucanus ibericus | 0.029–0.039 | 0.034–0.035 | 0.028–0.037 | 0.174 | 0.132–0.151 | 0.174 | 0.141–0.168 | NA | ||||
| Lucanus tetraodon | 0.125–0.129 | 0.124–0.128 | 0.122–0.130 | 0.168–0.179 | 0.098–0.123 | 0.151–0.156 | 0.132–0.151 | 0.131–0.136 | 0.024 | |||
| Lucanus (Pseudolucanus) macrophyllus | 0–0.012 | 0–0.014 | 0.006–0.015 | 0.159 | 0.116–0.141 | 0.147 | 0.130–0.145 | 0.028 | 0.120–0.124 | NA | ||
| Lucanus (Pseudolucanus) barbarossa | 0.153–0.163 | 0.155–0.161 | 0.155–0.167 | 0.004–0.058 | 0.127–0.171 | 0.153–0.165 | 0.146–0.167 | 0.166–0.172 | 0.159–0.177 | 0.149–0.157 | 0.053 | |
| unknown Lucanus sp. | 0.091–0.162 | 0.093–0.159 | 0.95–0.168 | 0.143–0.172 | 0.119–0.150 | 0.016–0.066 | 0.088–0.113 | 0.109–0.169 | 0.120–0.152 | 0.087–0.147 | 0.136–0.170 | 0.002–0.054 |
Nucleotide composition and K2P-distances at each codon position of the 670 bp COI region.
| Codon position | |||
|---|---|---|---|
| 1st | 2nd | 3rd | |
| % A | 31.4 | 18.9 | 38.2 |
| % T | 26.6 | 42.5 | 36.6 |
| % G | 25.6 | 16.2 | 4.9 |
| % C | 16.4 | 22.4 | 20.4 |
| K2P-distance | 0–0.107 | 0–0.032 | 0–0.999 |
These results do not show a distinct barcoding gap or other threshold to distinguish putative species, which is chiefly due to a lack of phylogenetic resolution to differentiate the said species and subspecies. If we consider the taxa of the Lucanus cervus cervus clade to be members of the same species, 99.4% of all intra(sub)specific comparisons showed K2P-distances below 5% and 99.8% of the pairwise inter(sub)specific distances were above 5%. Nucleotide diagnostics are listed in Appendix 2. No diagnostic combination of nucleotide positions and characters were found for the taxa of the Lucanus cervus cervus clade, Lucanus ibericus not included. As the number of species and the sample size per species are rather limited, the nucleotide diagnostics should be considered with caution.
The present study shows that the sequenced COI fragment could discriminate several of the investigated western Palaearctic Lucanus species and alleged subspecies of Lucanus cervus. Well differentiated species and subspecies were Lucanus cervus akbesianus, Lucanus cervus laticornis and Lucanus tetraodon, as well as the two eastern Asian species Lucanus formosanus and Lucanus hermani. Difficulties in molecular identification remained between Lucanus cervus fabiani and Lucanus (Pseudolucanus) barbarossa, Lucanus cervus judaicus and the unidentified Lucanus species, and between taxa of the Lucanus cervus cervus clade. Although thoroughly sampled within their distribution range, Lucanus cervus cervus and Lucanus cervus turcicus could not be discriminated with a barcoding approach. Likewise, three out of four samples of Lucanus cervus pentaphyllus possessed the most common haplotype of Lucanus cervus cervus. Next to introgression following recent or past hybridisation events, incomplete sorting of ancestral variation may be the reason for the polyphyletic pattern. It is not known if Lucanus can be infected with the endosymbiotic bacteria Wolbachia, which can cause mitochondrial introgression between closely related species (e.g.
Like Lucanus (Pseudolucanus) macrophyllus, one sample of Lucanus (Pseudolucanus) barbarossa, SB6, was embedded in the Lucanus cervus cervus clade, opposed to the other two samples that clustered with Lucanus cervus fabiani. The taxa of the latter group showed K2P-distances between 0.004 and 0.058, which indicates a close relationship between Lucanus cervus fabiani and Lucanus (Pseudolucanus) barbarossa, as well as Lucanus (Pseudolucanus) barbarossa being very variable. High intraspecific variability could be indicative of cryptic diversity or population structure (Diptera:
Finally, the Lucanus samples from Israel and Lebanon that were unidentified at the species level, seemed closely related and formed a paraphyletic clade with Lucanus cervus judaicus. Nevertheless, some of these samples could well be of a different species, indicated by the higher pairwise genetic distances (0.042–0.066). A detailed morphological and phylogenetic study is required here to investigate the number of species and relationship with Lucanus cervus judaicus.
A distinct barcoding gap was absent for several species and subspecies of Lucanus. This may either represent a low phylogenetic signal from the COI fragment for some relationships, a problem of basing a taxonomy on just one or a few morphological traits, or both. The use of the COI gene for barcoding purposes has had mixed results. High intraspecific variability (
This study revealed that while the 3’ terminus of COI contained sufficient information to resolve relationships among a number of closely related taxa, many others could not be robustly discriminated. Genotyping of additional specimens, especially of Lucanus (Pseudolucanus) macrophyllus, Lucanus ibericus, Lucanus cervus judaicus, Lucanus cervus fabiani and Lucanus cervus laticornis, as well as all western Palaearctic taxa is needed to fully explore COI genetic diversity and to investigate the roles of phenotypic plasticity, hybridisation and incomplete lineage sorting underlying stag beetle biodiversity and inform taxonomic investigations. We therefore see this study as a starting point for future research which should also endeavour to combine analysis of nuclear markers, such as the internal transcribed spacer (ITS) and 28S rRNA gene (e.g.
The work presented here was carried out in collaboration between all authors. AT, KDG, GA, PA and LB defined the subject and the design of the study. KDG designed methods and experiments in the laboratory and supervised laboratory work. KC analysed the data, interpreted results and wrote the paper. AT was responsible for collecting the samples and co-wrote the taxonomical part of the paper. JM discussed analyses. GA, ES, NMcK and PS provided five sequences and revised primarily the material and methods section and the interpretation of the results. MZ, LB and PA provided samples and co-wrote the paper, particularly the taxonomical section. DH and RM provided samples. All authors have contributed to, revised and approved the manuscript.
We want to thank the following people for generously providing tissue samples of stag beetles or information: E. Atay, M. Avci, L. Barbiero, R. Bekchiev, G. Bonamie, S. Boucher, C. Bouget, H. Brustel, D. G. Carrilero, L.R. Castro, G. De Coninck, J. Ibero Caballero, A.M. Cimaz, I. de las Monjas, M. Fremlin, N. Gouix, C. Hawes, J-P. Huang, N. Jansson, A. Kairouz, S. Korneyev, V.A. Korneyev, D. Kovalchuk, I. López Pérez, Á. Martínez García, M. Méndez, M. Murat, L. Nádai, I. Nel, E. Orbach, H. Podskalská, S. Rastrero Sánchez, S. Reicher, O. Rittner, F. Roviralta Peña, P. Šípek, L. Valladares, J.T. Smit, Á. R. Quirós Menéndez. Also many thanks to Leen Verschaeve, Nancy Van Liefferinge, An Van Breusegem, David Halfmaerten and Sabrina Neyrinck (INBO) for laboratory assistance. We appreciate the constructive comments of three anonymous reviewers.
Consensus Bayesian tree of 60 haplotypes of the 3’ end of the COI gene. Values given by the nodes are posterior probabilities above 0.70. (doi: 10.3897/zookeys.365.5526.app1) File format: Adobe PDF file (pdf).
Nucleotide diagnostics for (sub)species or species groups according to the Neighbour-Joining and Bayesian Inference tree topology. (doi: 10.3897/zookeys.365.5526.app2) File format: Adobe PDF file (pdf).