(C) 2011 Jing Chen. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
The taxonomic position of Hormaphis similibetulae Qiao & Zhang, 2004 has been reexamined. The phylogenetic position of Hormaphis similibetulae was inferred by maximum parsimony, maximum likelihood and Bayesian analyses on the basis of partial nuclear elongation factor-1α and mitochondrial tRNA leucine/cytochrome oxidase II sequences. The results showed that this species fell into the clade of Hamamelistes species, occupying a basal position, and was clearly distinct from other Hormaphis species. A closer relationship between Hormaphis similibetulae and Hamamelistes species was also revealed by life cycle analysis. Therefore, we conclude that Hormaphis similibetulae should be transferred to the genus Hamamelistes as Hamamelistes similibetulae (Qiao & Zhang), comb. n.
Hormaphidinae, Hamamelistes similibetulae, molecular evidence, biological evidence, new combination, China
The aphid tribe Hormaphidini in subfamily Hormaphidinae (Hemiptera: Aphididae) consists of three genera, Hamamelistes, Hormaphis and Protohormaphis (
The samples used in this study and the corresponding collection information are listed in Table 1. Eight species of Hormaphidini, covering all the species of Hamamelistes and Hormaphis were used as ingroups. Three species of Nipponaphidini were chosen as outgroups because Nipponaphidini is considered the sister group of Hormaphidini based on biological and phylogenetic data (
Total genomic DNA was extracted from single aphids preserved in 95% or 100% ethanol using a CTAB protocol modified from
Phylogenetic reconstructions were conducted by maximum
parsimony (MP), maximum likelihood (ML) and Bayesian analyses for each
single gene and a combined dataset. The partition homogeneity test (
Collection information and GenBank accession numbers for aphid samples used in this study.
| Species | Host | Locality | Date | Voucher | EF-1× | tRNA/COII |
|---|---|---|---|---|---|---|
| Hamamelistes betulinus (Horvath) | Betula davurica | Japan: Yamanashi, Masutomi | 17 Jul. 1998 | 98081 | AF454599* | AF328782* |
| Hamamelis japonica | Japan: Aomori, Temmabayashi | 7 Aug. 1998 | 98132 | AF454596* | AF328775* | |
| Betula platyphylla | Japan: Tokyo, Okutamako | 20 May 1999 | 99121 | AF454597* | AF328780* | |
| Betula platyphylla | Japan: Hokkaido, Sapporo | 15 Jun. 1999 | 99187 | AF454598* | AF328781* | |
| Hamamelistes kagamii (Monzen) | Hamamelis japonica | Japan: Yamanashi, Masutomi | 17 Jul. 1998 | 98084 | AF454600* | AF328772* |
| Betula grossa | Japan: Yamanashi, Sanjonoyu | 20 May 1999 | 99118 | AF454601* | AF328779* | |
| Hamamelis japonica | Japan: Saitama, Shomaru Pass | 8 Jul. 1999 | 99209 | AF454603* | AF328773* | |
| Hamamelis japonica | Japan: Saitama, Shomaru Pass | 8 Jul. 1999 | 99220 | AF454602* | AF328774* | |
| Hamamelistes miyabeiHamamelistes miyabei (Matsumura) | Hamamelis japonica | Japan: Yamanashi, Masutomi | 17 Jul. 1998 | 98086 | AF454595* | AF328771* |
| Betula maximowicziana | Japan: Hokkaido, Sapporo | 5 Sep. 1998 | 98151 | AF454593* | AF328776* | |
| Betula maximowicziana | Japan: Gumma, Mt. Akagi | 25 May 1999 | 99146 | AF454594* | AF328777* | |
| Betula maximowicziana | Japan: Hokkaido, Sapporo | 15 Jun. 1999 | 99182 | AF454592* | AF328778* | |
| Hamamelistes spinosus Shimer | Hamamelis japonica | USA: Washington, DC | May 1993 | 93-23 | AF454606* | AF328783* |
| Betula nigra | USA: UT, Logan | 28 May 1999 | 99-54 | AF454607* | AF454619* | |
| Betula nigra | USA: WI, Madison | 28 Jun. 1999 | 99-57 | AF454608* | None | |
| Hormaphis betulae (Mordvilko) | Betula platyphylla | Japan: Yamanashi, Masutomi | 17 Jul. 1998 | 98078 | AF454609* | None |
| Hamamelis japonica | Japan: Saitama, Shomaru Pass | 21 May 1999 | 99130 | AF454610* | AF454622* | |
| Betula platyphylla | Japan: Tokyo, Kazahari Pass | 26 Jul. 1999 | 99224 | AF454611* | AF454623* | |
| Betula sp. | China: Jilin, Ji’an | 13 Aug. 2004 | 15214 | DQ493864* | JF730745 | |
| Hormaphis cornu (Shimer) | Hamamelis virginiana | USA: Georgia, Athens | 8 Jun. 1994 | 94-93 | AF454612* | AF454621* |
| Hormaphis hamamelidis (Fitch) | Hamamelis virginiana | USA: Connecticut, Danielson | 1 Aug. 1998 | 98-05 | AF454613* | AF454620* |
| Hormaphis similibetulae Qiao & Zhang | Betula albosinensis | China: Tibet, Gongbo’gyamda | 5 Jul. 2002 | 13549 | DQ493849* | JF730746 |
| Betula albosinensis | China: Tibet, Linzhi | 6 Aug. 2003 | 15318 | DQ493866* | JF730747 | |
| Neohormaphis wuyiensis Qiao & Jiang | Quercus sp. | China: Fujian, Mt. Wuyi | 18 Jul. 2003 | 14525 | DQ493858* | JF730748 |
| Nipponaphis distyliicola Monzen | Quercus glauca | Japan: Shinkiba, Tokyo | 16 Apr. 1999 | 99008 | AF454614* | AF454626* |
| Thoracaphis quercifoliae Ghosh | Quercus sp. | China: Fujian, Mt. Wuyi | 18 Jul. 2003 | 14526_2 | DQ493851* | JF730749 |
* Sequences from GenBank.
The final alignments of EF-1α (excluding three introns) and tRNA/COII sequences consisted of 826 and 761 sites, with 131 and 165 parsimony-informative sites, respectively. A single 1- to 2-base-long indel was found in the tRNA. The genetic distance between two distinct samples of Hormaphis similibetulae was 0 for EF-1α and 0.001 for tRNA/COII. The distances of both genes between Hormaphis similibetulae and Hamamelistes species were much smaller than those between Hormaphis similibetulae and the other Hormaphis species (EF-1α: average of 0.040 and range of 0.038–0.042 to Hamamelistes, average of 0.082 and range of 0.078–0.092 to Hormaphis; tRNA/COII: average of 0.080 and range of 0.071–0.085 to Hamamelistes, average of 0.106 and range of 0.102–0.112 to Hormaphis).
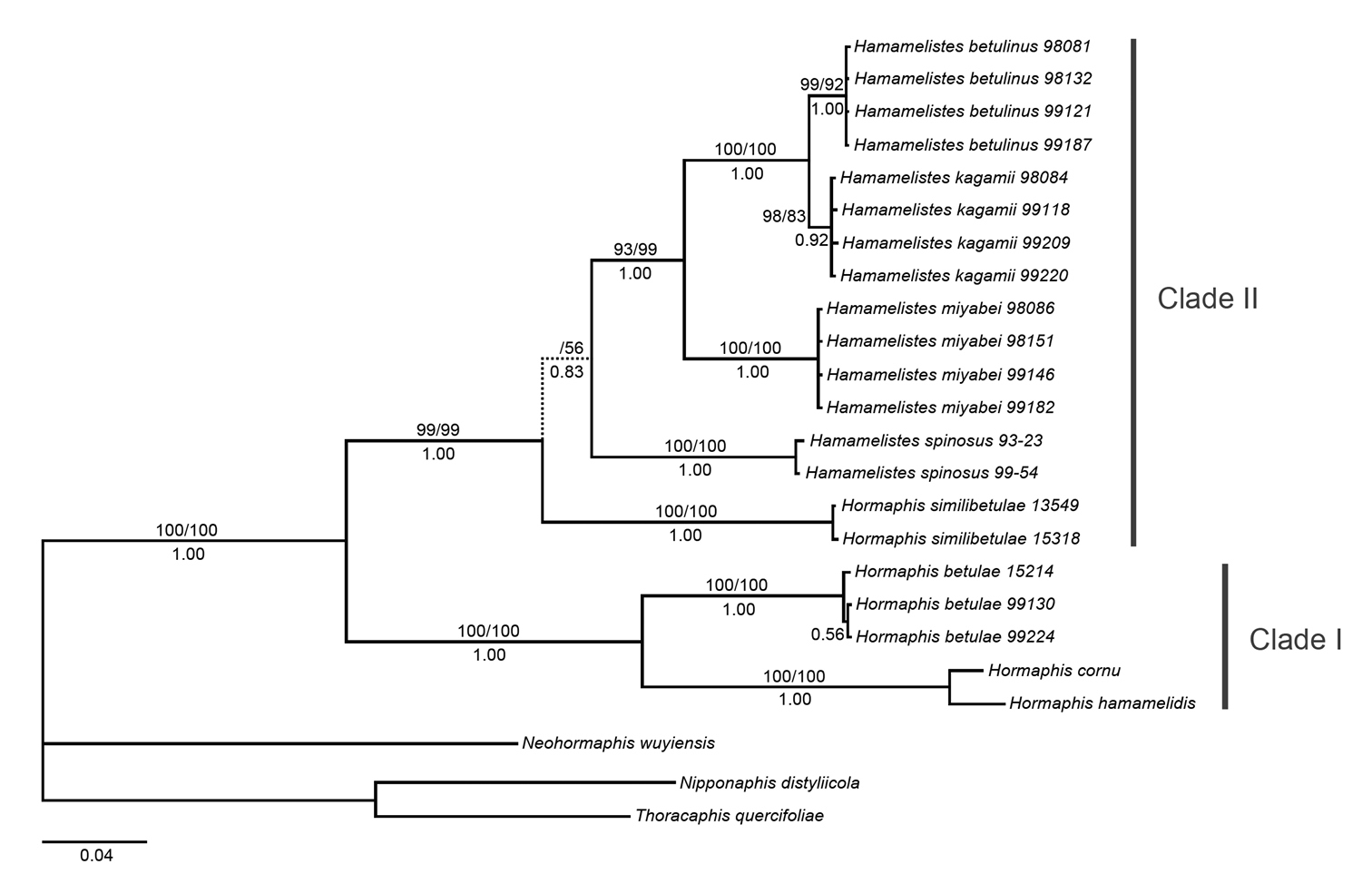
For phylogenetic analyses, the partition homogeneity test found no significant conflict between EF-1α and mtDNA (P=0.05), indicating that information from both genes could be combined. Combined analysis resulted in similar topology to that obtained in single gene analyses and with higher support for most nodes, so only the combined dataset results were presented. MP analysis yielded eight most parsimonious trees with a length of 611 steps (CI=0.705401, RI=0.845626). ML analysis produced one ML tree based on the optimal model GTR+G selected by AIC in Modeltest 3.7. The 50% majority-rule consensus tree inferred from Bayesian analysis is shown in Fig. 1 and resulted in a topology essentially identical to that obtained in ML analysis, but was different from the strict consensus of MP trees in the position of Hormaphis similibetulae. All ingroup taxa constituted a monophyletic group with respect to these outgroups and formed two clades. Clade I (100% MP BS, 100% ML BS, 1.00 PP) was comprised of Hormaphis betulae, Hormaphis cornu, and Hormaphis hamamelidis. Clade II (99% MP BS, 99% ML BS, 1.00 PP) consisted of all the Hamamelistes species and Hormaphis similibetulae. Within clade II, two distinct samples of Hormaphis similibetulae clustered together (100% MP BS, 100% ML BS, 1.00 PP) and were placed as the outermost branch in ML and Bayesian analyses, just as the results based on EF-1α. However, MP analysis revealed the same topology as the mitochondrial analysis: Hormaphis similibetulae and Hamamelistes spinosus were sister groups, although the support value was low (53% BS), and together formed the basal lineage within clade II.
Phylogenetic tree reconstructed from the combined dataset of EF-1α and tRNA/COII sequences. The Bayesian topology and branch lengths are shown. Values above the branches are MP and ML bootstrap percentages, respectively, and Bayesian posterior probabilities are shown below the branches. The broken line indicates inconsistent branch.
The results of genetic distances and phylogenetic analyses strongly suggested that Hormaphis similibetulae was more closely related to Hamamelistes than to Hormaphis. Hormaphis similibetulae was distinguished by its unique biology, forming galls on leaves of Betula. Because of the high morphological similarity with Hormaphis betulae (Mordvilko), it was placed under the genus Hormaphis (
The phylogenetic position of Hormaphis similibetulae was inferred by MP, ML and Bayesian analyses on the basis of nuclear EF-1α and mitochondrial tRNA/COII sequences. In all phylogenetic analyses, Hormaphis similibetulae clustered firmly with Hamamelistes and was placed as a basal lineage, clearly differed from other Hormaphis species. Life cycle similarities also indicated that Hormaphis similibetulae was more closely related to Hamamelistes species. We therefore conclude that Hormaphis similibetulae should be transferred to the genus Hamamelistes as Hamamelistes similibetulae (Qiao & Zhang), comb. n.
Thanks are due to X. L. Huang, G. X. Qiao and N. Qiao for their collections. The work was supported by the National Natural Sciences Foundation of China (Nos. 30830017, 31061160186), National Science Funds for Distinguished Young Scientists (No. 31025024), National Science Fund for Fostering Talents in Basic Research (No. J0930004), and a grant (No. O529YX5105) from the Key Laboratory of the Zoological Systematics and Evolution of the Chinese Academy of Sciences.